| Entry |
Thumbnail Image |
Name |
Description |
Object |
Legend |
|
hsa04070
|

|
Phosphatidylinositol signaling system |
|
...5332 (PLCB4), 5333 (PLCD1), 5335 (PLCG1), 5336 (PLCG2), 84812 (PLCD4), 89869 (PLCZ1) 113026 (PLCD3),... |
PTEN 2.7.1.158 PI(3,4)P PI(3,5)P PI(5)P PLC PLC 5PP-IP Focal adhesion I(1,3,4)P I(1,3,4,5)P I(4)P I(... |
|
hsa04625
|

|
C-type lectin receptor signaling pathway |
C-type lectin receptors (CLRs) are a large superfamily of proteins characterized by the presence of ... |
...), 6237 (RRAS) 5580 (PRKCD) 4792 (NFKBIA) 5336 (PLCG2) 5594 (MAPK1), 5595 (MAPK3) 4772 (NFATC1), 477... |
C-TYPE LECTIN RECEPTOR SIGNALING PATHWAY Dectin-1 Dectin-2 Mincle MCL FcRγ DC-SIGN LSP-1 KSR1 CN... |
|
hsa04650
|

|
Natural killer cell mediated cytotoxicity |
Natural killer (NK) cells are lymphocytes of the innate immune system that are involved in early def... |
...459 (IFNGR1), 3460 (IFNGR2) 5335 (PLCG1), 5336 (PLCG2) 27040 (LAT) 5551 (PRF1) 3133 (HLA-E) 3458 (IF... |
IgG NFAT CaN PI(3,4,5)P Fyn TRAIL IFNγR PLCγ LAT Perforin HLA-E IFN-γ GM-CSF TNF-α PKC 3BP2 PLCγ LAT... |
|
hsa05206
|

|
MicroRNAs in cancer |
MicroRNA (miRNA) is a cluster of small non-encoding RNA molecules of 21 - 23 nucleotides in length, ... |
... 2261 (FGFR3) 5156 (PDGFRA) 5335 (PLCG1), 5336 (PLCG2) 5894 (RAF1) 3845 (KRAS) 5578 (PRKCA), 5579 (P... |
MicroRNAs IN CANCER Lung cancer Lung epithelial cell Tumorigenesis Survival Invasion / metastasi... |
|
hsa05214
|

|
Glioma |
Gliomas are the most common of the primary brain tumors and account for more than 40% of all central... |
...), 1870 (E2F2), 1871 (E2F3) 5335 (PLCG1), 5336 (PLCG2) 163688 (CALML6), 51806 (CALML5), 801 (CALM1),... |
EGF TGFα PDGF IGF-1 EGFR PDGFR IGFR EGF TGFα PDGF IGF-1 EGFR PDGFR IGFR CAM CAMK PKC Shc Grb2 Sos Ra... |
|
hsa00562
|

|
Inositol phosphate metabolism |
|
...5332 (PLCB4), 5333 (PLCD1), 5335 (PLCG1), 5336 (PLCG2), 84812 (PLCD4), 89869 (PLCZ1), 9651 (PLCH2) 6... |
1-Phosphatidyl-1D-myo-inositol-5P 2.7.1.149 1D-myo-Inositol- 1,4,5,6P 3.1.3.36 2.7.1.151 2.7.1.151 3... |
|
hsa01100
|

|
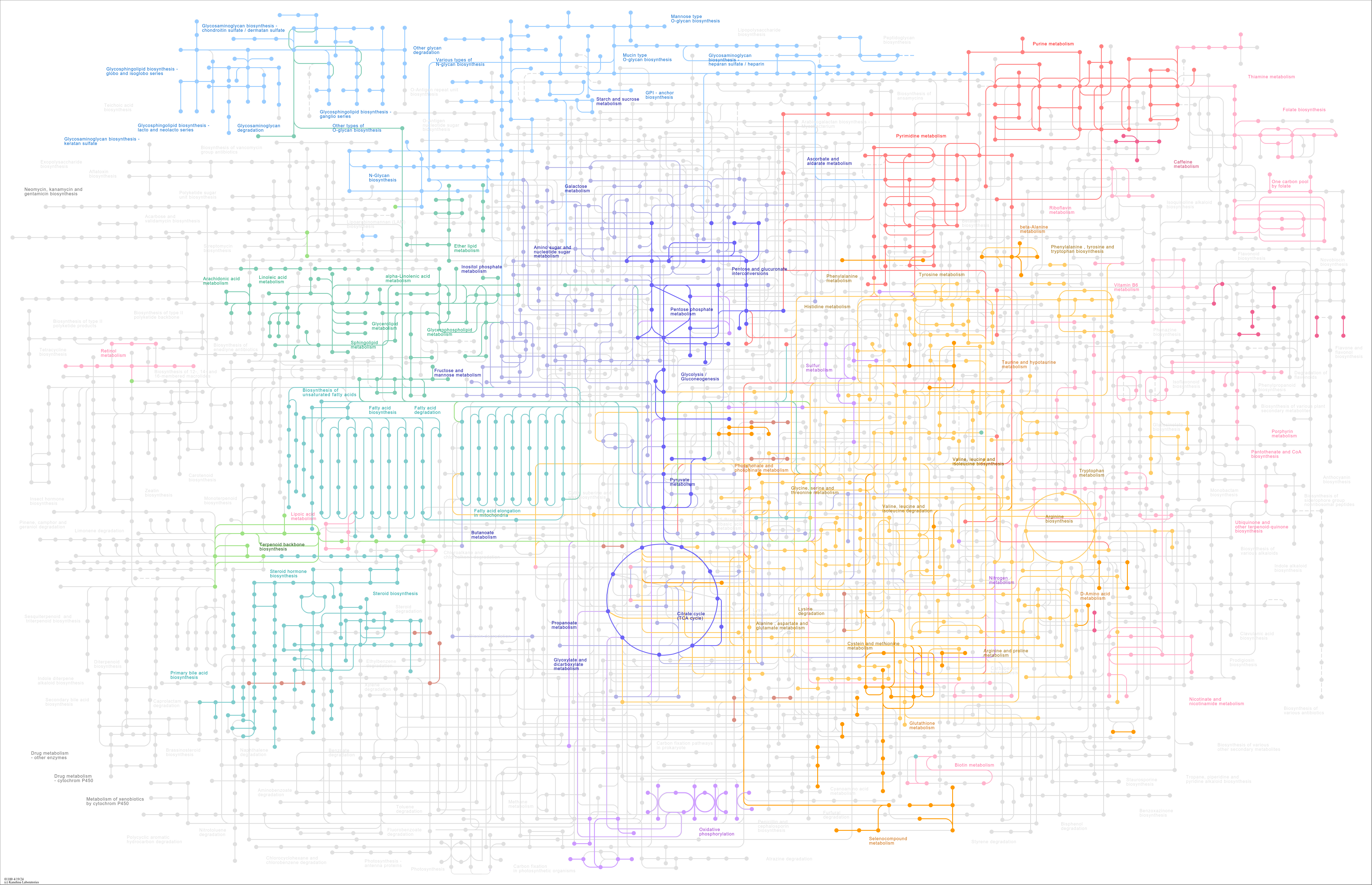
Metabolic pathways |
|
...5332 (PLCB4), 5333 (PLCD1), 5335 (PLCG1), 5336 (PLCG2), 84812 (PLCD4), 89869 (PLCZ1), 9651 (PLCH2) 2... |
Monoterpenoid biosynthesis One carbon pool by folate Flavonoid biosynthesis Starch and sucrose metab... |
|
hsa01521
|

|
EGFR tyrosine kinase inhibitor resistance |
EGFR is a tyrosine kinase that participates in the regulation of cellular homeostasis. EGFR also ser... |
... 5579 (PRKCB), 5582 (PRKCG) 5335 (PLCG1), 5336 (PLCG2) 5290 (PIK3CA), 5291 (PIK3CB), 5293 (PIK3CD), ... |
Non-small cell lung cancer Her3 Her2 EGFR Pancreatic cancer mTOR signaling pathway GSK-3 eI... |
|
hsa04012
|

|
ErbB signaling pathway |
The ErbB family of receptor tyrosine kinases (RTKs) couples binding of extracellular growth factor l... |
... 817 (CAMK2D), 818 (CAMK2G) 5335 (PLCG1), 5336 (PLCG2) 1839 (HBEGF) 3084 (NRG1) 2069 (EREG) 685 (BTC... |
Non-small cell lung cancer Grb2 Elk ErbB-4 ErbB-4 ErbB-3 ErbB-3 STAT5 ErbB-2 ErbB-2 ErbB-2 ErbB-2... |
|
hsa04014
|

|
Ras signaling pathway |
The Ras proteins are GTPases that function as molecular switches for signaling pathways regulating c... |
...), 94235 (GNG8) 3363 (HTR7) 5335 (PLCG1), 5336 (PLCG2) 7535 (ZAP70) 27040 (LAT) 10298 (PAK4), 106821... |
MAPK signaling pathway RAS SIGNALING PATHWAY ERK MEK Raf-1 Ras PI3K-Akt signaling pathway Grb2 SOS... |
|
hsa04020
|

|
Calcium signaling pathway |
Ca2+ that enters the cell from the outside is a principal source of signal Ca2+. Entry of Ca2+ is dr... |
... 5027 (P2RX7), 9127 (P2RX6) 5335 (PLCG1), 5336 (PLCG2) 56848 (SPHK2), 8877 (SPHK1) 89869 (PLCZ1) 340... |
Long term depression Long term potentiation Phosphatidylinositol signaling pathway Apoptosis ... |
|
hsa04062
|

|
Chemokine signaling pathway |
Inflammatory immune response requires the recruitment of leukocytes to the site of inflammation upon... |
...5331 (PLCB3), 5332 (PLCB4), 5335 (PLCG1), 5336 (PLCG2) 25759 (SHC2), 399694 (SHC4), 53358 (SHC3), 64... |
Migration Apoptosis Degranulation Cellular shape changes Cell survival chemokineR chemokine JAK2/3 ... |
|
hsa04064
|

|
NF-kappa B signaling pathway |
Nuclear factor-kappa B (NF-kappa B) is the generic name of a family of transcription factors that fu... |
...U) 598 (BCL2L1) 598 (BCL2L1) 597 (BCL2A1) 5336 (PLCG2) 6850 (SYK) 4067 (LYN) 695 (BTK) 3932 (LCK) 29... |
Cytokine-cytokine receptor interaction Bcl-XL Bcl-2 c-IAP1/2 IκBα IRAK1/4 MyD88 TRAF2/5 RIP1 TRADD I... |
|
hsa04066
|
|
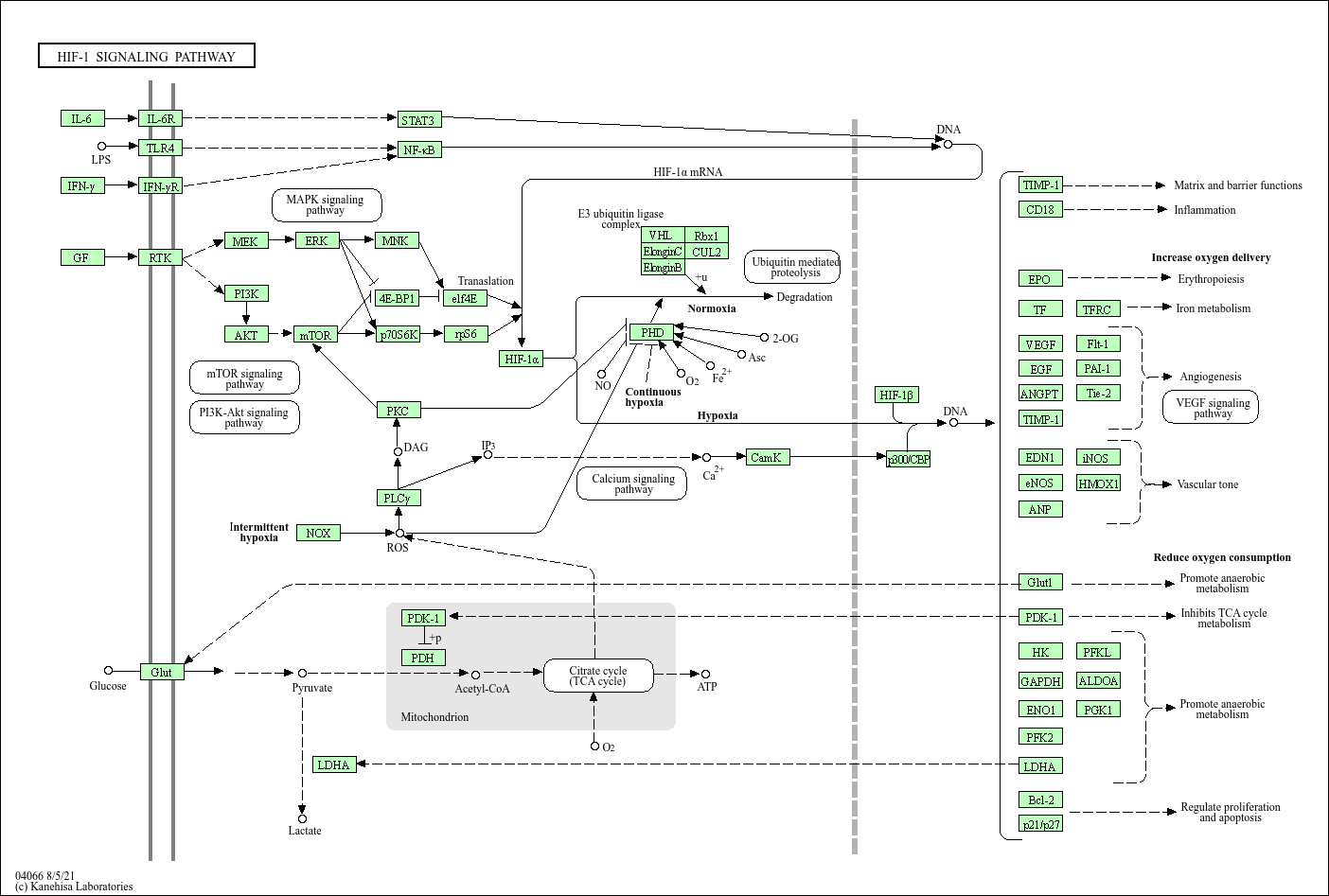
HIF-1 signaling pathway |
Hypoxia-inducible factor 1 (HIF-1) is a transcription factor that functions as a master regulator of... |
...T2) 2475 (MTOR) 1536 (CYBB) 5335 (PLCG1), 5336 (PLCG2) 5578 (PRKCA), 5579 (PRKCB), 5582 (PRKCG) 6513... |
CUL2 Rbx1 ElonginB ElonginC VEGF PHD VHL HIF-1β p300/CBP HIF-1α Glut1 HIF-1 SIGNALING PATHWAY mTOR... |
|
hsa04072
|

|
Phospholipase D signaling pathway |
Phospholipase D (PLD) is an essential enzyme responsible for the production of the lipid second mess... |
... 5737 (PTGFR), 9170 (LPAR2) 5335 (PLCG1), 5336 (PLCG2) 10411 (RAPGEF3), 11069 (RAPGEF4) 5578 (PRKCA)... |
MAPK signaling pathway PHOSPHOLIPASE D SIGNALING PATHWAY ERK MEK Raf-1 Ras PI3K-Akt signaling pat... |
|
hsa04360
|

|
Axon guidance |
Axon guidance represents a key stage in the formation of neuronal network. Axons are guided by a var... |
... 7224 (TRPC5), 7225 (TRPC6) 5335 (PLCG1), 5336 (PLCG2) 8440 (NCK2) 10298 (PAK4), 106821730 (BUB1B-PA... |
Par6 Par3 CXCR4 ERK Rac Rac Rac DCC NFAT CaN Netrin-G1 Netrin-4 Slit3 Slit2 Slit1 FAK Cdc42 ROCK PAK... |
|
hsa04370
|

|
VEGF signaling pathway |
There is now much evidence that VEGFR-2 is the major mediator of VEGF-driven responses in endothelia... |
... I2) 998 (CDC42) 3791 (KDR) 5335 (PLCG1), 5336 (PLCG2) 56848 (SPHK2), 8877 (SPHK1) 5743 (PTGS2) 1001... |
Cdc42 DAG Calcium signaling pathway Focal adhesion MAPK signaling pathway VEGFR2 PLCγ SPK PIP PGI CO... |
|
hsa04380
|

|
Osteoclast differentiation |
The osteoclasts, multinucleared cells originating from the hematopoietic monocyte-macrophage lineage... |
...TEC) 6850 (SYK) 29760 (BLNK), 3937 (LCP2) 5336 (PLCG2) 5530 (PPP3CA), 5532 (PPP3CB), 5533 (PPP3CC), ... |
OSTEOCLAST DIFFERENTIATION c-Fms TRAF2/6 RANK RANKL M-CSF OPG CYLD Gab2 NIK IKKα IKKγ IKKα IKKβ IκB... |
|
hsa04611
|

|
Platelet activation |
Platelets play a key and beneficial role for primary hemostasis on the disruption of the integrity o... |
...P6) 3673 (ITGA2), 3688 (ITGB1) 2814 (GP5) 5336 (PLCG2) 10125 (RASGRP1), 10235 (RASGRP2) 5906 (RAP1A)... |
PLATELET ACTIVATION cAMP DAG Calcium signaling pathway Platelet PAR1/4 P2Y1 ADP TXA2 PLCβ GP VI α2... |
|
hsa04613
|

|
Neutrophil extracellular trap formation |
Neutrophils play a central role in innate immune defense. One of the mechanisms of neutrophil action... |
...5331 (PLCB3), 5332 (PLCB4), 5335 (PLCG1), 5336 (PLCG2) 5578 (PRKCA), 5579 (PRKCB), 5582 (PRKCG) 5894... |
Neutrophil FcγRIIA FcγRI Src Syk IgG PI3K PIP PLC DAG PKC Raf1 MEK ERK1/2 Akt NOD-like receptor sign... |