| Entry |
|
|---|
| Name |
Starch and sucrose metabolism - Actinomyces capricornis |
|---|
| Class |
Metabolism; Carbohydrate metabolism
|
|---|
| Pathway map |
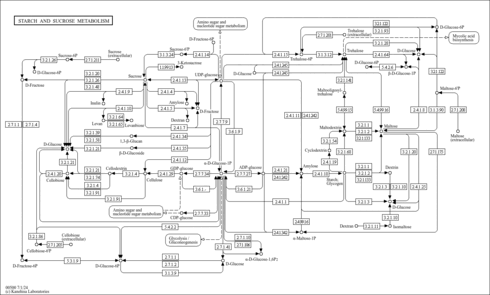
|
|---|
| Other DBs |
|
|---|
| Organism |
Actinomyces capricornis [GN: acap]
|
|---|
| Gene |
|
|---|
| Compound |
| C00663 | beta-D-Glucose 1-phosphate |
| C00689 | alpha,alpha'-Trehalose 6-phosphate |
| C01231 | alpha-D-Glucose 1,6-bisphosphate |
| C03323 | (2,1-beta-D-Fructosyl)n |
| C04534 | 6-Phospho-beta-D-glucosyl-(1,4)-D-glucose |
| C06400 | 1-alpha-D-[(1->4)-alpha-D-Glucosyl](n-1)-alpha-D-glucopyranoside |
| C20237 | alpha-Maltose 1-phosphate |
|
|---|
Related
pathway |
| acap00520 | Amino sugar and nucleotide sugar metabolism |
|
|---|
| KO pathway |
|
|---|
| LinkDB |
|
|---|