 | | PATHWAY: blr01100 | |
| Entry |
|
|---|
| Name |
Metabolic pathways - Brevibacillus laterosporus |
|---|
| Class |
|
|---|
| Pathway map |
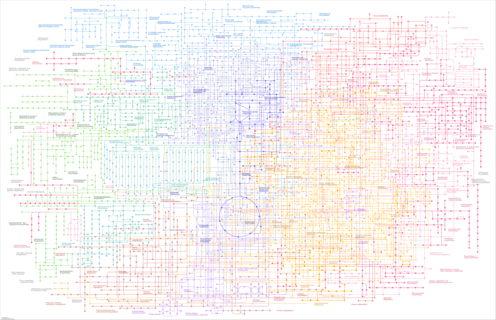
|
|---|
| Module |
| blr_M00007 | Pentose phosphate pathway, non-oxidative phase, fructose 6P => ribose 5P [PATH:blr01100] |
| blr_M00010 | Citrate cycle, first carbon oxidation, oxaloacetate => 2-oxoglutarate [PATH:blr01100] |
| blr_M00011 | Citrate cycle, second carbon oxidation, 2-oxoglutarate => oxaloacetate [PATH:blr01100] |
| blr_M00019 | Valine/isoleucine biosynthesis, pyruvate => valine / 2-oxobutanoate => isoleucine [PATH:blr01100] |
| blr_M00022 | Shikimate pathway, phosphoenolpyruvate + erythrose-4P => chorismate [PATH:blr01100] |
| blr_M00045 | Histidine degradation, histidine => N-formiminoglutamate => glutamate [PATH:blr01100] |
| blr_M00053 | Deoxyribonucleotide biosynthesis, ADP/GDP/CDP/UDP => dATP/dGTP/dCTP/dUTP [PATH:blr01100] |
| blr_M00133 | Polyamine biosynthesis, arginine => agmatine => putrescine => spermidine [PATH:blr01100] |
| blr_M00570 | Isoleucine biosynthesis, threonine => 2-oxobutanoate => isoleucine [PATH:blr01100] |
| blr_M00572 | Pimeloyl-ACP biosynthesis, BioC-BioH pathway, malonyl-ACP => pimeloyl-ACP [PATH:blr01100] |
| blr_M00579 | Phosphate acetyltransferase-acetate kinase pathway, acetyl-CoA => acetate [PATH:blr01100] |
| blr_M00883 | Lipoic acid biosynthesis, animals and bacteria, octanoyl-ACP => dihydrolipoyl-H => dihydrolipoyl-E2 [PATH:blr01100] |
| blr_M00909 | UDP-N-acetyl-D-glucosamine biosynthesis, prokaryotes, glucose => UDP-GlcNAc [PATH:blr01100] |
| blr_M00916 | Pyridoxal-P biosynthesis, R5P + glyceraldehyde-3P + glutamine => pyridoxal-P [PATH:blr01100] |
| blr_M00924 | Cobalamin biosynthesis, anaerobic, uroporphyrinogen III => sirohydrochlorin => cobyrinate a,c-diamide [PATH:blr01100] |
|
|---|
| Organism |
Brevibacillus laterosporus [GN: blr]
|
|---|
Related
pathway |
| blr00040 | Pentose and glucuronate interconversions |
| blr00051 | Fructose and mannose metabolism |
| blr00053 | Ascorbate and aldarate metabolism |
| blr00130 | Ubiquinone and other terpenoid-quinone biosynthesis |
| blr00250 | Alanine, aspartate and glutamate metabolism |
| blr00260 | Glycine, serine and threonine metabolism |
| blr00270 | Cysteine and methionine metabolism |
| blr00280 | Valine, leucine and isoleucine degradation |
| blr00290 | Valine, leucine and isoleucine biosynthesis |
| blr00330 | Arginine and proline metabolism |
| blr00361 | Chlorocyclohexane and chlorobenzene degradation |
| blr00400 | Phenylalanine, tyrosine and tryptophan biosynthesis |
| blr00430 | Taurine and hypotaurine metabolism |
| blr00440 | Phosphonate and phosphinate metabolism |
| blr00520 | Amino sugar and nucleotide sugar metabolism |
| blr00523 | Polyketide sugar unit biosynthesis |
| blr00525 | Acarbose and validamycin biosynthesis |
| blr00541 | O-Antigen nucleotide sugar biosynthesis |
| blr00542 | O-Antigen repeat unit biosynthesis |
| blr00592 | alpha-Linolenic acid metabolism |
| blr00625 | Chloroalkane and chloroalkene degradation |
| blr00630 | Glyoxylate and dicarboxylate metabolism |
| blr00660 | C5-Branched dibasic acid metabolism |
| blr00710 | Carbon fixation by Calvin cycle |
| blr00760 | Nicotinate and nicotinamide metabolism |
| blr00770 | Pantothenate and CoA biosynthesis |
| blr00900 | Terpenoid backbone biosynthesis |
| blr00907 | Pinene, camphor and geraniol degradation |
| blr00997 | Biosynthesis of various other secondary metabolites |
| blr00998 | Biosynthesis of various antibiotics |
| blr00999 | Biosynthesis of various plant secondary metabolites |
| blr01053 | Biosynthesis of siderophore group nonribosomal peptides |
|
|---|
| KO pathway |
|
|---|
| LinkDB |
|
|---|
|
|