 | | PATHWAY: ege01110 | |
| Entry |
|
|---|
| Name |
Biosynthesis of secondary metabolites - Duffyella gerundensis |
|---|
| Class |
|
|---|
| Pathway map |
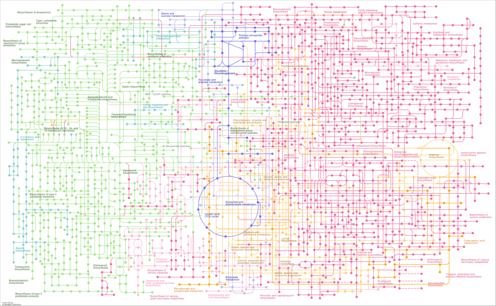
| ege01110 | Biosynthesis of secondary metabolites |
|
|---|
| Module |
| ege_M00019 | Valine/isoleucine biosynthesis, pyruvate => valine / 2-oxobutanoate => isoleucine [PATH:ege01110] |
| ege_M00022 | Shikimate pathway, phosphoenolpyruvate + erythrose-4P => chorismate [PATH:ege01110] |
| ege_M00024 | Phenylalanine biosynthesis, chorismate => phenylpyruvate => phenylalanine [PATH:ege01110] |
| ege_M00117 | Ubiquinone biosynthesis, prokaryotes, chorismate (+ polyprenyl-PP) => ubiquinol [PATH:ege01110] |
| ege_M00125 | Riboflavin biosynthesis, plants and bacteria, GTP => riboflavin/FMN/FAD [PATH:ege01110] |
|
|---|
| Organism |
Duffyella gerundensis [GN: ege]
|
|---|
Related
pathway |
| ege00053 | Ascorbate and aldarate metabolism |
| ege00130 | Ubiquinone and other terpenoid-quinone biosynthesis |
| ege00250 | Alanine, aspartate and glutamate metabolism |
| ege00260 | Glycine, serine and threonine metabolism |
| ege00270 | Cysteine and methionine metabolism |
| ege00280 | Valine, leucine and isoleucine degradation |
| ege00290 | Valine, leucine and isoleucine biosynthesis |
| ege00400 | Phenylalanine, tyrosine and tryptophan biosynthesis |
| ege00440 | Phosphonate and phosphinate metabolism |
| ege00523 | Polyketide sugar unit biosynthesis |
| ege00525 | Acarbose and validamycin biosynthesis |
| ege00592 | alpha-Linolenic acid metabolism |
| ege00630 | Glyoxylate and dicarboxylate metabolism |
| ege00770 | Pantothenate and CoA biosynthesis |
| ege00900 | Terpenoid backbone biosynthesis |
| ege00997 | Biosynthesis of various other secondary metabolites |
| ege00999 | Biosynthesis of various plant secondary metabolites |
| ege01053 | Biosynthesis of siderophore group nonribosomal peptides |
|
|---|
| KO pathway |
|
|---|
| LinkDB |
|
|---|
|
|