 | | PATHWAY: rgl01100 | |
| Entry |
|
|---|
| Name |
Metabolic pathways - Rhodanobacter glycinis |
|---|
| Class |
|
|---|
| Pathway map |
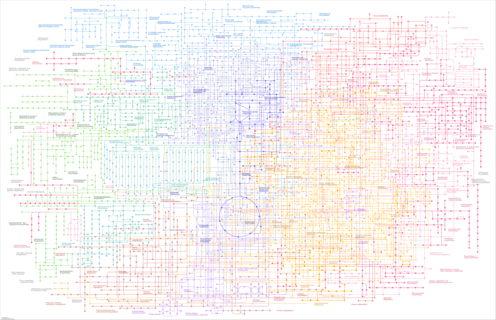
|
|---|
| Module |
| rgl_M00006 | Pentose phosphate pathway, oxidative phase, glucose 6P => ribulose 5P [PATH:rgl01100] |
| rgl_M00007 | Pentose phosphate pathway, non-oxidative phase, fructose 6P => ribose 5P [PATH:rgl01100] |
| rgl_M00010 | Citrate cycle, first carbon oxidation, oxaloacetate => 2-oxoglutarate [PATH:rgl01100] |
| rgl_M00011 | Citrate cycle, second carbon oxidation, 2-oxoglutarate => oxaloacetate [PATH:rgl01100] |
| rgl_M00024 | Phenylalanine biosynthesis, chorismate => phenylpyruvate => phenylalanine [PATH:rgl01100] |
| rgl_M00045 | Histidine degradation, histidine => N-formiminoglutamate => glutamate [PATH:rgl01100] |
| rgl_M00053 | Deoxyribonucleotide biosynthesis, ADP/GDP/CDP/UDP => dATP/dGTP/dCTP/dUTP [PATH:rgl01100] |
| rgl_M00125 | Riboflavin biosynthesis, plants and bacteria, GTP => riboflavin/FMN/FAD [PATH:rgl01100] |
| rgl_M00552 | D-galactonate degradation, De Ley-Doudoroff pathway, D-galactonate => glycerate-3P [PATH:rgl01100] |
| rgl_M00845 | Arginine biosynthesis, glutamate => acetylcitrulline => arginine [PATH:rgl01100] |
| rgl_M00909 | UDP-N-acetyl-D-glucosamine biosynthesis, prokaryotes, glucose => UDP-GlcNAc [PATH:rgl01100] |
| rgl_M00948 | Hydroxyproline degradation, trans-4-hydroxy-L-proline => 2-oxoglutarate [PATH:rgl01100] |
|
|---|
| Organism |
Rhodanobacter glycinis [GN: rgl]
|
|---|
Related
pathway |
| rgl00040 | Pentose and glucuronate interconversions |
| rgl00051 | Fructose and mannose metabolism |
| rgl00053 | Ascorbate and aldarate metabolism |
| rgl00121 | Secondary bile acid biosynthesis |
| rgl00130 | Ubiquinone and other terpenoid-quinone biosynthesis |
| rgl00250 | Alanine, aspartate and glutamate metabolism |
| rgl00260 | Glycine, serine and threonine metabolism |
| rgl00270 | Cysteine and methionine metabolism |
| rgl00280 | Valine, leucine and isoleucine degradation |
| rgl00290 | Valine, leucine and isoleucine biosynthesis |
| rgl00311 | Penicillin and cephalosporin biosynthesis |
| rgl00330 | Arginine and proline metabolism |
| rgl00361 | Chlorocyclohexane and chlorobenzene degradation |
| rgl00400 | Phenylalanine, tyrosine and tryptophan biosynthesis |
| rgl00430 | Taurine and hypotaurine metabolism |
| rgl00520 | Amino sugar and nucleotide sugar metabolism |
| rgl00523 | Polyketide sugar unit biosynthesis |
| rgl00525 | Acarbose and validamycin biosynthesis |
| rgl00540 | Lipopolysaccharide biosynthesis |
| rgl00541 | O-Antigen nucleotide sugar biosynthesis |
| rgl00592 | alpha-Linolenic acid metabolism |
| rgl00625 | Chloroalkane and chloroalkene degradation |
| rgl00630 | Glyoxylate and dicarboxylate metabolism |
| rgl00660 | C5-Branched dibasic acid metabolism |
| rgl00760 | Nicotinate and nicotinamide metabolism |
| rgl00770 | Pantothenate and CoA biosynthesis |
| rgl00900 | Terpenoid backbone biosynthesis |
| rgl00907 | Pinene, camphor and geraniol degradation |
| rgl00999 | Biosynthesis of various plant secondary metabolites |
| rgl01040 | Biosynthesis of unsaturated fatty acids |
|
|---|
| KO pathway |
|
|---|
| LinkDB |
|
|---|
|
|