| Entry |
|
|---|
| Name |
Galactose metabolism - Mucilaginibacter mali |
|---|
| Class |
Metabolism; Carbohydrate metabolism
|
|---|
| Pathway map |
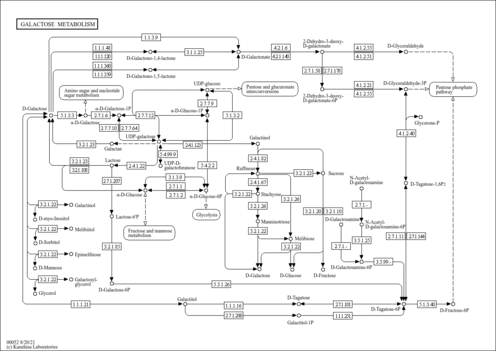
|
|---|
| Other DBs |
|
|---|
| Organism |
Mucilaginibacter mali [GN: mmab]
|
|---|
| Gene |
|
|---|
| Compound |
| C00118 | D-Glyceraldehyde 3-phosphate |
| C00446 | alpha-D-Galactose 1-phosphate |
| C00668 | alpha-D-Glucose 6-phosphate |
| C01113 | D-Galactose 6-phosphate |
| C01132 | N-Acetyl-D-galactosamine |
| C01216 | 2-Dehydro-3-deoxy-D-galactonate |
| C01235 | alpha-D-Galactosyl-(1->3)-1D-myo-inositol |
| C01286 | 2-Dehydro-3-deoxy-6-phospho-D-galactonate |
| C02669 | D-Galactono-1,5-lactone |
| C03383 | D-Galactono-1,4-lactone |
| C03733 | UDP-alpha-D-galactofuranose |
| C03785 | D-Tagatose 1,6-bisphosphate |
| C05401 | 3-beta-D-Galactosyl-sn-glycerol |
| C05404 | D-Gal alpha 1->6D-Gal alpha 1->6D-Glucose |
| C06376 | N-Acetyl-D-galactosamine 6-phosphate |
| C06377 | D-Galactosamine 6-phosphate |
|
|---|
Related
pathway |
| mmab00040 | Pentose and glucuronate interconversions |
| mmab00520 | Amino sugar and nucleotide sugar metabolism |
|
|---|
| KO pathway |
|
|---|