| Entry |
|
|---|
| Name |
Riboflavin metabolism - Olea europaea var. sylvestris (wild olive) |
|---|
| Class |
Metabolism; Metabolism of cofactors and vitamins
|
|---|
| Pathway map |
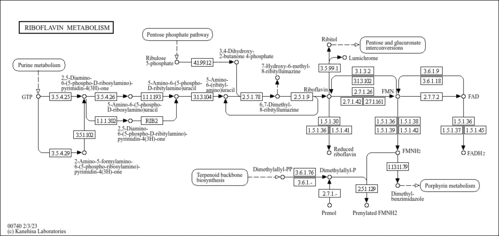
|
|---|
| Module |
| oeu_M00125 | Riboflavin biosynthesis, plants and bacteria, GTP => riboflavin/FMN/FAD [PATH:oeu00740] |
|
|---|
| Other DBs |
|
|---|
| Organism |
Olea europaea var. sylvestris (wild olive) [GN: oeu]
|
|---|
| Gene |
|
|---|
| Compound |
| C00235 | Dimethylallyl diphosphate |
| C01268 | 5-Amino-6-(5'-phosphoribosylamino)uracil |
| C01304 | 2,5-Diamino-6-(5-phospho-D-ribosylamino)pyrimidin-4(3H)-one |
| C04332 | 6,7-Dimethyl-8-(D-ribityl)lumazine |
| C04454 | 5-Amino-6-(5'-phospho-D-ribitylamino)uracil |
| C04732 | 5-Amino-6-(1-D-ribitylamino)uracil |
| C05995 | 7-Hydroxy-6-methyl-8-ribityllumazine |
| C15556 | L-3,4-Dihydroxybutan-2-one 4-phosphate |
| C15563 | 2-Amino-5-formylamino-6-(5-phospho-D-ribosylamino)pyrimidin-4(3H)-one |
| C18910 | 2,5-Diamino-6-(5-phospho-D-ribitylamino)pyrimidin-4(3H)-one |
| C21214 | Dimethylallyl phosphate |
|
|---|
| Reference |
|
|---|
| Authors |
Vitreschak AG, Rodionov DA, Mironov AA, Gelfand MS. |
|---|
| Title |
Regulation of riboflavin biosynthesis and transport genes in bacteria by transcriptional and translational attenuation. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Fischer M, Haase I, Feicht R, Richter G, Gerhardt S, Changeux JP, Huber R, Bacher A. |
|---|
| Title |
Biosynthesis of riboflavin: 6,7-dimethyl-8-ribityllumazine synthase of Schizosaccharomyces pombe. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Haase I, Fischer M, Bacher A, Schramek N. |
|---|
| Title |
Temperature-dependent presteady state kinetics of lumazine synthase from the hyperthermophilic eubacterium Aquifex aeolicus. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Fischer M, Romisch W, Illarionov B, Eisenreich W, Bacher A. |
|---|
| Title |
Structures and reaction mechanisms of riboflavin synthases of eubacterial and archaeal origin. |
|---|
| Journal |
|
|---|
Related
pathway |
| oeu00040 | Pentose and glucuronate interconversions |
| oeu00900 | Terpenoid backbone biosynthesis |
|
|---|
| KO pathway |
|
|---|
| LinkDB |
|
|---|