 | | PATHWAY: salp01100 | |
| Entry |
|
|---|
| Name |
Metabolic pathways - Salvelinus sp. IW2-2015 (Arctic char) |
|---|
| Class |
|
|---|
| Pathway map |
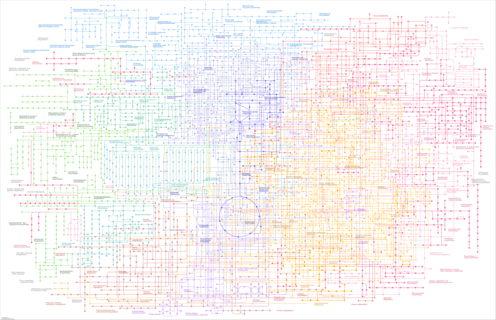
|
|---|
| Module |
| salp_M00007 | Pentose phosphate pathway, non-oxidative phase, fructose 6P => ribose 5P [PATH:salp01100] |
| salp_M00042 | Catecholamine biosynthesis, tyrosine => dopamine => noradrenaline => adrenaline [PATH:salp01100] |
| salp_M00046 | Pyrimidine degradation, uracil => beta-alanine, thymine => 3-aminoisobutanoate [PATH:salp01100] |
| salp_M00053 | Deoxyribonucleotide biosynthesis, ADP/GDP/CDP/UDP => dATP/dGTP/dCTP/dUTP [PATH:salp01100] |
| salp_M00107 | Steroid hormone biosynthesis, cholesterol => pregnenolone => progesterone [PATH:salp01100] |
| salp_M00110 | C19/C18-Steroid hormone biosynthesis, pregnenolone => androstenedione => estrone [PATH:salp01100] |
| salp_M00128 | Ubiquinone biosynthesis, eukaryotes, 4-hydroxybenzoate + polyprenyl-PP => ubiquinone [PATH:salp01100] |
| salp_M00130 | Inositol phosphate metabolism, PI=> PIP2 => Ins(1,4,5)P3 => Ins(1,3,4,5)P4 [PATH:salp01100] |
| salp_M00131 | Inositol phosphate metabolism, Ins(1,3,4,5)P4 => Ins(1,3,4)P3 => myo-inositol [PATH:salp01100] |
| salp_M00143 | NADH dehydrogenase (ubiquinone) Fe-S protein/flavoprotein complex, mitochondria [PATH:salp01100] |
| salp_M00955 | CMP-Neu5Ac/Neu5Gc biosynthesis, animals, UDP-GlcNAc => CMP-Neu5Ac => CMP-Neu5Gc [PATH:salp01100] |
| salp_M00976 | C19-Steroid hormone biosynthesis, pregnenolone => testosterone => dihydrotestosterone [PATH:salp01100] |
|
|---|
| Organism |
Salvelinus sp. IW2-2015 (Arctic char) [GN: salp]
|
|---|
Related
pathway |
| salp00040 | Pentose and glucuronate interconversions |
| salp00130 | Ubiquinone and other terpenoid-quinone biosynthesis |
| salp00250 | Alanine, aspartate and glutamate metabolism |
| salp00260 | Glycine, serine and threonine metabolism |
| salp00280 | Valine, leucine and isoleucine degradation |
| salp00290 | Valine, leucine and isoleucine biosynthesis |
| salp00400 | Phenylalanine, tyrosine and tryptophan biosynthesis |
| salp00440 | Phosphonate and phosphinate metabolism |
| salp00513 | Various types of N-glycan biosynthesis |
| salp00514 | Other types of O-glycan biosynthesis |
| salp00520 | Amino sugar and nucleotide sugar metabolism |
| salp00524 | Neomycin, kanamycin and gentamicin biosynthesis |
| salp00532 | Glycosaminoglycan biosynthesis - chondroitin sulfate / dermatan sulfate |
| salp00533 | Glycosaminoglycan biosynthesis - keratan sulfate |
| salp00534 | Glycosaminoglycan biosynthesis - heparan sulfate / heparin |
| salp00541 | Biosynthesis of various nucleotide sugars |
| salp00563 | Glycosylphosphatidylinositol (GPI)-anchor biosynthesis |
| salp00601 | Glycosphingolipid biosynthesis - lacto and neolacto series |
| salp00603 | Glycosphingolipid biosynthesis - globo and isoglobo series |
| salp00604 | Glycosphingolipid biosynthesis - ganglio series |
| salp00630 | Glyoxylate and dicarboxylate metabolism |
| salp00760 | Nicotinate and nicotinamide metabolism |
| salp00980 | Metabolism of xenobiotics by cytochrome P450 |
| salp01040 | Biosynthesis of unsaturated fatty acids |
|
|---|
| KO pathway |
|
|---|
|
|