| Entry |
Thumbnail |
Name |
Description |
Object |
Legend |
|
hsa04152
|

|
AMPK signaling pathway
|
AMP-activated protein kinase (AMPK) is a serine threonine kinase that is highly conserved through evolution. AMPK system acts as a sensor of cellular energy status. It is activated by increases in the ...
|
C00031 (D-Glucose)
C05981 (Phosphatidylinositol-3,4,5-trisphosphate)
C00008 (ADP)
C00020 (AMP)
C00003 (NAD+)
C00076 (Calcium cation)
C00024 (Acetyl-CoA)
C00083 (Malonyl-CoA)
C00040 (Acyl-CoA)
C00085 (D-Fructose ...
|
AMPK
INS
Glucose
INSR
Akt
P13K
PDK1/2
IRS
PIP
AMPK SIGNALING PATHWAY
Glycolysis / Gluconeogenesis
DNA
PI3K-Akt signaling pathway
IGF1
IGF1R
FOXO
Cell cycle
Insulin signaling pathway
GLUT4
LKB1
ADP
AMP
Low ...
|
|
hsa04210
|

|
Apoptosis
|
Apoptosis is a genetically programmed process for the elimination of damaged or redundant cells by activation of caspases (aspartate-specific cysteine proteases). The onset of apoptosis is controlled by ...
|
C00076 (Calcium cation)
C05981 (Phosphatidylinositol-3,4,5-trisphosphate)
C00319 (Sphingosine)
C00027 (Hydrogen peroxide)
598 (BCL2L1), D09935 (Navitoclax (JAN/USAN/INN)), D09936 (Navitoclax dihydrochloride ...
|
Mitochondrion
p53 signaling pathway
TNF signaling pathway
Bcl-XL
ICAD
ATM
p53
ENDO-G
AIF
Bcl-2
IAP/XIAP
CAD
CASP9
CASP7
CASP3
Bad
NF-κB
IAP/XIAP
CASP6
IκBα
Apaf-1
CytC
Bid
CASP8
CASP10
Calpain
Akt/PKB
NIK
IKK
FLIP
PI3K
TRAF2
RIP1
TRADD
FADD
TRADD
FADD
IL-3R
TrkA
TNF-R1
TRAIL-R
Fas
IL-3
NGF
TNFα
APOPTOSIS
TRAIL
Fas-L
Survival ...
|
|
hsa04211
|

|
Longevity regulating pathway
|
Regulation of longevity depends on genetic and environmental factors. Caloric restriction (CR), that is limiting food intake, is recognized in mammals as the best characterized and most reproducible strategy ...
|
C05981 (Phosphatidylinositol-3,4,5-trisphosphate)
C00575 (3',5'-Cyclic AMP)
C03582 (Resveratrol)
C07909 (Sirolimus)
C07151 (Metformin)
C00003 (NAD+)
C00020 (AMP)
C00076 (Calcium cation)
2308 (FOXO1), 2309 ...
|
FOXO
S6K
mTOR
Akt
P13K
PIP
LONGEVITY REGULATING PATHWAY
DNA
IGF-I
IGF-IR
IRS1-4
Ras
cAMP
PKA
AMPK
INS
INSR
Insulin signaling pathway
mTOR signaling pathway
PI3K-Akt signaling pathway
CREB
FOXO signaling ...
|
|
hsa04213
|

|
Longevity regulating pathway - multiple species
|
Aging is a complex process of accumulation of molecular, cellular, and organ damage, leading to loss of function and increased vulnerability to disease and death. Despite the complexity of aging, recent ...
|
C05981 (Phosphatidylinositol-3,4,5-trisphosphate)
C05981 (Phosphatidylinositol-3,4,5-trisphosphate)
C05981 (Phosphatidylinositol-3,4,5-trisphosphate)
C00575 (3',5'-Cyclic AMP)
C00575 (3',5'-Cyclic AMP)
2308 ...
|
FOXO
S6K
mTOR
Akt
P13K
PIP
LONGEVITY REGULATING PATHWAY - MULTIPLE SPECIES
DNA
IGF-I
IGF-IR
IRS1-4
Ras
Mammals
Flies
Worms
Yeast
dFOXO
S6K
mTOR
Akt
P13K
PIP
DNA
DILP
InR
CHICO
DAF-16
Akt
AGE-1
PIP
DNA
DAF-2
Gpr1
Ras2
PKA
Msn2/4
RSK-1
mTOR
Sch9
mTOR
RIM15
4E-BP
PHA-4
Gis1
INS
IST-1
Nutrients ...
|
|
hsa04215
|

|
Apoptosis - multiple species
|
Apoptosis is an evolutionarily conserved process used by multicellular organisms to developmentally regulate cell number or to eliminate cells that are potentially detrimental to the organism. The major ...
|
79444 (BIRC7)
840 (CASP7)
317 (APAF1)
666 (BOK)
54205 (CYCS), H00978 (Thrombocytopenia (THC))
842 (CASP9), D11195 (Rimiducid (USAN/INN))
840 (CASP7)
836 (CASP3)
329 (BIRC2), 330 (BIRC3), 331 (XIAP), 332 ...
|
APOPTOSIS - MULTIPLE SPECIES
Mitochondrion
DIAP1
DRONC
DRICE
GRIM
RPR
HID
dArk
Dcp-1
DEBCL
RHG family
CytC
Drosophila
Mammals
Casp9
Casp7
Casp3
XIAP
APAF-1
SMAC
HtrA2
ARTS
Apoptosis
Apoptosis
C. elegans
Egl-1
CED-4
CED-3
Apoptosis
Apoptosis ...
|
|
hsa04216
|

|
Ferroptosis
|
Ferroptosis is a regulated form of cell death and characterized by a production of reactive oxygen species (ROS) from accumulated iron and lipid peroxidation. It can be induced by experimental compounds ...
|
C00025 (L-Glutamate)
C00491 (L-Cystine)
C00051 (Glutathione)
C21478 (Erastin)
C00097 (L-Cysteine)
C00669 (gamma-L-Glutamyl-L-cysteine)
C00418 ((R)-Mevalonate)
C11378 (Ubiquinone-10)
C14819 (Fe3+)
C14818 ...
|
FERROPTOSIS
SLC7A11
Glutamate
Cystine
p53
GPX4
GSH
Mitochondria
VDAC2/3
Erastin
Cysteine
GCL
γGC
SLC3A2
MVA
CoQ
TFR1
Endosome
Erastin
Fenton reaction
TFR1
STEAP3
TFR1
GSSG
GSS
PCBP1/2
Ferroportin
PCBP2
Ferritin
ATG5/7
Ferritin
ZIP8/14
DMT1
Autolyosome
Iron ...
|
|
hsa04217
|

|
Necroptosis
|
Necroptosis is a programmed form of necrosis. It can be initiated by different stimuli, such as tumor necrosis factor (TNF), TNF-related apoptosis-inducing ligand (TRAIL), Fas ligand (FasL), interferon ...
|
C00076 (Calcium cation)
C00305 (Magnesium cation)
C00002 (ATP)
C00027 (Hydrogen peroxide), C16844 (Hydroxyl radical)
C00002 (ATP)
C00195 (N-Acylsphingosine)
C00319 (Sphingosine)
C00076 (Calcium cation)
C00076 ...
|
NECROPTOSIS
TNF
TNFR1
TRADD
TRAF2/5
MLKL
TLR4
IFNR
LPS
IFN
JAKs
TRAILR
TRAIL
Fas
FasL
IAPs
Casp8
cFLIP
MLKL
CYLD
TRPM7
Release
Endosome
TLR3
dsRNA
RIPK1
FADD
A20
Pore formation
FADD
RIPK1
RIPK1
Membrane ...
|
|
hsa04218
|

|
Cellular senescence
|
Cellular senescence is a state of irreversible cellular arrest and can be triggered by a number of factors, such as telomere shortening, oncogene activation, irradiation, DNA damage and oxidative stress ...
|
C16844 (Hydroxyl radical), C00027 (Hydrogen peroxide)
C07909 (Sirolimus)
C00076 (Calcium cation)
C16844 (Hydroxyl radical), C00027 (Hydrogen peroxide)
C00076 (Calcium cation)
C00076 (Calcium cation)
C05981 ...
|
CELLULAR SENESCENCE
NORE1A
PP1A
C-Myc
HIPK2
ARF
MDM2
p53
P53 signaling pathway
PI3K
AKT
TSC1/2
RHEB
mTOR
FOXO3
SIRT1
Ras
MEK
Ets1
Raf
CDK4/6
CycD
ERK
p16
p21
CDK2
CycE
E2F
p38
p53
ROS
MKK3/6
CDK2
CycA
B-MYB
FOXM1
MuvB
SIRT1
ATM
CHK2
MRE11
RAD50
NBS1
GADD45
ATR
CDC25A
CHK1
RAD9
RAD1
HUS1
CDK1
CycB
CDK2
CycE
CDK4/6
CycD
Cellular ...
|
|
hsa04260
|

|
Cardiac muscle contraction
|
Contraction of the heart is a complex process initiated by the electrical excitation of cardiac myocytes (excitation-contraction coupling, ECC). In cardiac myocytes, Ca2+ influx induced by activation of ...
|
C00076 (Calcium cation)
C00076 (Calcium cation)
C00076 (Calcium cation)
C00076 (Calcium cation)
C01330 (Sodium cation)
C01330 (Sodium cation)
C00076 (Calcium cation)
C00080 (H+)
C00080 (H+)
C01330 (Sodium ...
|
CARDIAC MUSCLE CONTRACTION
T-tubule
Sarcoplasmic Reticulum (SR)
DHPR
RyR2
NCX
NCX
Systole
Diastole
SERCA2a
TnC
3Na
3Na
Myofilaments
TnI
TnC
TnT
Actin
TPM
Myosin
Sarcolemma
Cardiac myocyte
Calcium signaling ...
|
|
hsa04261
|

|
Adrenergic signaling in cardiomyocytes
|
Cardiac myocytes express at least six subtypes of adrenergic receptor (AR) which include three subtypes of beta-AR (beta-1, beta-2, beta-3) and three subtypes of the alpha-1-AR (alpha-1A, alpha-1B, and ...
|
C00076 (Calcium cation)
C00076 (Calcium cation)
C00076 (Calcium cation)
C01330 (Sodium cation)
C01330 (Sodium cation)
C00076 (Calcium cation)
C00238 (Potassium cation)
C01330 (Sodium cation)
C00080 (H+)
C01330 ...
|
ADRENERGIC SIGNALING IN CARDIOMYOCYTES
T-tubule
Sarcoplasmic Reticulum (SR)
DHPR
RyR2
NCX
NCX
SERCA2a
TnC
3Na
3Na
Myofilaments
TnT
TnI
Actin
TPM
Myosin
Sarcolemma
Cardiac myocyte
Calcium signaling pathway
Crossbridge
NaK
3Na
Depolarization
NHE
β1AR
β2AR
PKA
PI3K
Akt
cAMP
DNA
ERK
CREB
PLB
I-1
PP1
PP2A
CaM
CaMKII
AT-IIR
PLC
PKCα
Cardiac ...
|
|
hsa04270
|

|
Vascular smooth muscle contraction
|
The vascular smooth muscle cell (VSMC) is a highly specialized cell whose principal function is contraction. On contraction, VSMCs shorten, thereby decreasing the diameter of a blood vessel to regulate ...
|
C01245 (D-myo-Inositol 1,4,5-trisphosphate)
C00165 (Diacylglycerol)
C00076 (Calcium cation)
C00219 (Arachidonate)
C14748 (20-HETE)
C00238 (Potassium cation)
C00533 (Nitric oxide)
C00575 (3',5'-Cyclic AMP)
C00942 ...
|
Calcium signaling pathway
MLCK
MLC
MLCP
ROCK
PKC
RhoA
RhoGEF
VASCULAR SMOOTH MUSCLE CONTRACTION
Actin
Relaxation
ADRA1
Gαq /11
12/13
PLC
DAG
Sarcoplasmic reticulum (SR)
CaM
VOC
Contraction
CPI-17
MLCP
MLC
Actin
Arachidonic ...
|
|
hsa04310
|

|
Wnt signaling pathway
|
Wnt proteins are secreted morphogens that are required for basic developmental processes, such as cell-fate specification, progenitor-cell proliferation and the control of asymmetric cell division, in ...
|
C00076 (Calcium cation)
4088 (SMAD3), H00800 (Loeys-Dietz syndrome)
4772 (NFATC1), 4773 (NFATC2), 4775 (NFATC3), 4776 (NFATC4), H02664 (Joint contracture, osteochondromas, and B-cell lymphoma)
5578 (PRKCA) ...
|
p53 signaling pathway
Adherens junction
SMAD3
TGF-β signaling pathway
Cell cycle
WNT SIGNALING PATHWAY
MAPK signaling pathway
Focal adhesion
Ubiquitin medeated proteolysis
MAPK signaling pathway
Alzheimer's ...
|
|
hsa04330
|

|
Notch signaling pathway
|
The Notch signaling pathway is an evolutionarily conserved, intercellular signaling mechanism essential for proper embryonic development in all metazoan organisms in the Animal kingdom. The Notch proteins ...
|
C22528 (Notch intracellular domain)
23385 (NCSTN), H00681 (Acne inversa), D08869 (Begacestat (USAN/INN)), D09010 (Tarenflurbil (USAN/INN)), D09377 (Semagacestat (USAN/INN)), D09869 (Avagacestat (USAN)) ...
|
NCSTN
APH-1
PSEN
MAPK signaling pathway
HDAC
CIR
Gro/TLE
CtBP
Hairless
SMRT
PreTα
Hes1/5
CSL
SKIP
HATs
MAML
ADAM17
PSE2
Deltex
Numb
Serrate
Notch
Dvl
NOTCH SIGNALING PATHWAY
Fringe
Delta
γ-Secretase ...
|
|
hsa04340
|

|
Hedgehog signaling pathway
|
The Hedgehog (Hh) signaling pathway has numerous roles in the control of cell proliferation, tissue patterning, stem cell maintenance and development. The primary cilium is an important center for transduction ...
|
C00575 (3',5'-Cyclic AMP)
6608 (SMO), H00039 (Basal cell carcinoma), H01556 (Meningioma), H01667 (Medulloblastoma), D09992 (Vismodegib (USAN/INN)), D10119 (Sonidegib (USAN/INN)), D10324 (Patidegib (USAN/INN)) ...
|
Smo
Gpr161
cAMP
CK1
GSK3β
PKA
Gli
Kif7
GliR
Cilium
Cytoplasm
Without Hh
Vesicle
Cul1
β-TrCP
Nucleus
HEDGEHOG SIGNALING PATHWAY
(Gli repressor form)
Gli
Ptc
Hhip
DNA
Ptc
Smo
With Hh
Smo
GliA
Cilium
Cytoplasm
Gli
Kif7
Sufu
Ptc
co-R
Smurf1/2
Degradation
Ptc
Vesicle
EVC
GRK2
CK1
Nucleus
Gli
Ptc
Hhip
DNA
Smo
Vesicle
Gpr161
β-Arrestin
Kif3a
Degradation
Spop
Cul3
Sufu
Cul3
Spop
Degradation
(Gli ...
|
|
hsa04350
|

|
TGF-beta signaling pathway
|
The transforming growth factor-beta (TGF-beta) family members, which include TGF-betas, activins and bone morphogenetic proteins (BMPs), are structurally related secreted cytokines found in species ranging ...
|
C14819 (Fe3+)
4052 (LTBP1), H00557 (Cutis laxa)
7057 (THBS1)
8454 (CUL1)
9978 (RBX1)
4089 (SMAD4), H00019 (Pancreatic cancer), H00020 (Colorectal cancer), H00533 (Hereditary hemorrhagic telangiectasia) ...
|
LTBP1
THBS1
Cul1
Rbx1
TGF-BETA SIGNALING PATHWAY
Apoptosis
Cell Cycle
Ubiquitin mediated proteolysis
MAPK signaling pathway
Smad4
Smad4
NodalRII
NodalRI
Smad2/3
Nodal
Smad6/7
Smad2/3
Pitx2
SP1
p300
p15
c-Myc
DP1
E2F4/5
p107
ActivinRII
ActivinRI
Activin
FST
Lefty
p70S6K
PP2A
ROCK1
RhoA
Smad4
SARA
Smad2/3
Skp1
ERK
Smad4
Smad6/7
Smurf1/2
TGFβ ...
|
|
hsa04360
|

|
Axon guidance
|
Axon guidance represents a key stage in the formation of neuronal network. Axons are guided by a variety of guidance factors, such as netrins, ephrins, Slits, and semaphorins. These guidance cues are read ...
|
C00076 (Calcium cation)
C00076 (Calcium cation)
C00076 (Calcium cation)
C00076 (Calcium cation)
50855 (PARD6A), 84552 (PARD6G), 84612 (PARD6B)
56288 (PARD3)
2770 (GNAI1), 2771 (GNAI2), 2773 (GNAI3), H00033 ...
|
Par6
Par3
CXCR4
ERK
Rac
Rac
Rac
DCC
NFAT
CaN
Netrin-G1
Netrin-4
Slit3
Slit2
Slit1
FAK
Cdc42
ROCK
PAK
RhoA
RhoA
Regulation of actin cytoskeleton
Regulation of actin cytoskeleton
Cytokine-cytokine receptor ...
|
|
hsa04370
|

|
VEGF signaling pathway
|
There is now much evidence that VEGFR-2 is the major mediator of VEGF-driven responses in endothelial cells and it is considered to be a crucial signal transducer in both physiologic and pathologic angiogenesis ...
|
C00533 (Nitric oxide)
C00076 (Calcium cation)
C01245 (D-myo-Inositol 1,4,5-trisphosphate)
C00165 (Diacylglycerol)
C05981 (Phosphatidylinositol-3,4,5-trisphosphate)
C01312 (Prostaglandin I2)
998 (CDC42) ...
|
Cdc42
DAG
Calcium signaling pathway
Focal adhesion
MAPK signaling pathway
VEGFR2
PLCγ
SPK
PIP
PGI
COX2
cPLA2
HSP27
MAPKAPK
CALN
Akt/PKB
PI3K
NFAT
PKC
FAK
p38
Paxillin
VRAP
Src
Sck
VEGF
VEGF SIGNALING ...
|
|
hsa04371
|
|
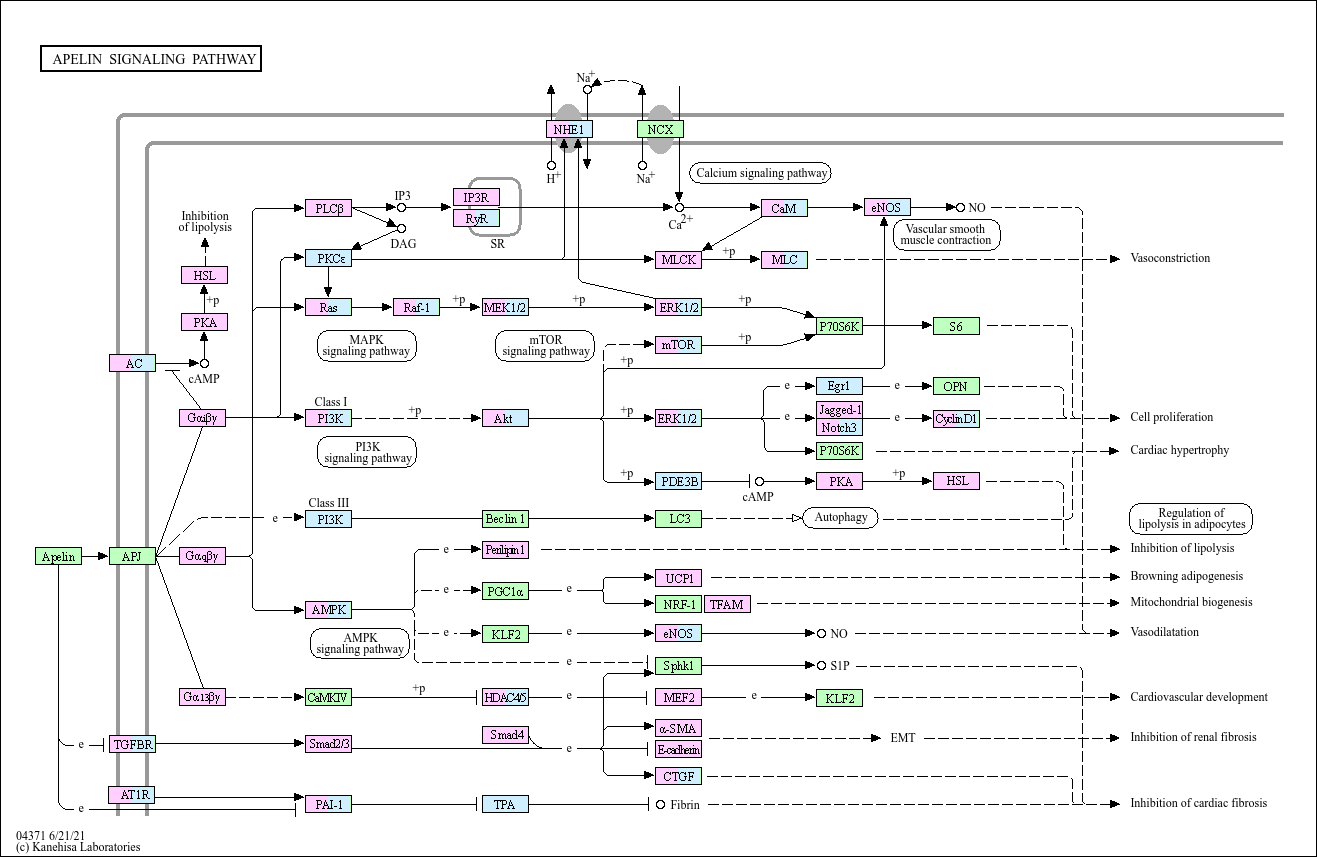
Apelin signaling pathway
|
Apelin is an endogenous peptide capable of binding the apelin receptor (APJ), which was originally described as an orphan G-protein-coupled receptor. Apelin and APJ are widely expressed in various tissues ...
|
C01245 (D-myo-Inositol 1,4,5-trisphosphate)
C00076 (Calcium cation)
C01330 (Sodium cation)
C00080 (H+)
C00165 (Diacylglycerol)
C01330 (Sodium cation)
C00575 (3',5'-Cyclic AMP)
C00575 (3',5'-Cyclic AMP)
C00533 ...
|
APELIN SIGNALING PATHWAY
APJ
AMPK
Sphk1
PAI-1
PI3K
Akt
MEK1/2
ERK1/2
P70S6K
PLCβ
IP3
IP3R
RyR
mTOR
Ras
Raf-1
KLF2
TGFBR
CTGF
PGC1α
TFAM
NRF-1
Mitochondrial biogenesis
MLCK
MLC
DAG
PKCε
NCX
Cyclin D1
Cell ...
|
|
hsa04380
|

|
Osteoclast differentiation
|
The osteoclasts, multinucleared cells originating from the hematopoietic monocyte-macrophage lineage, are responsible for bone resorption. Osteoclastogenesis is mainly regulated by signaling pathways activated ...
|
C01245 (D-myo-Inositol 1,4,5-trisphosphate)
C00076 (Calcium cation)
C00027 (Hydrogen peroxide)
1436 (CSF1R), H00028 (Choriocarcinoma), H01807 (Hereditary diffuse leukoencephalopathy with spheroids), D02664 ...
|
OSTEOCLAST DIFFERENTIATION
c-Fms
TRAF2/6
RANK
RANKL
M-CSF
OPG
CYLD
Gab2
NIK
IKKα
IKKγ
IKKα
IKKβ
IκB
NFκB
TAK1
TAB1/2
MKK6
MEK1
JNK
p38
NFATc1
IFNβ
DNA
IFNβ
IFNAR
Jak1/Tyk2
STAT1/2
IRF9
Src
PI3K
Akt
Cell ...
|
|
hsa04390
|

|
Hippo signaling pathway
|
Hippo signaling is an evolutionarily conserved signaling pathway that controls organ size from flies to humans. In humans and mice, the pathway consists of the MST1 and MST2 kinases, their cofactor Salvador ...
|
23286 (WWC1)
4771 (NF2), H00015 (Malignant pleural mesothelioma), H01438 (Neurofibromatosis type 2), H01556 (Meningioma), H01667 (Medulloblastoma)
122786 (FRMD6), 79981 (FRMD1)
60485 (SAV1)
6788 (STK3)
10413 ...
|
HIPPO SIGNALING PATHWAY
KIBRA
Mer
FRMD
SAV1
Mst1/2
YAP/TAZ
Lats1/2
Crb
α-Catenin
RASSF1A
CTGF
Birc5
AREG
DNA
PP1
ASPP2
14-3-3
Mob
YAP/TAZ
AMOT
Pals1
PATJ
Tight junction
Adherens junction
E-Cad
E-Cad
Lgl
Scrib
Dlg
TEAD
Ajub
Cellular ...
|
|
hsa04392
|

|
Hippo signaling pathway - multiple species
|
Hippo signaling pathways control diverse aspects of cell proliferation, survival, and morphogenesis in eukaryotes. The core organization of these networks is conserved over a billion years of evolution ...
|
23286 (WWC1)
4771 (NF2), H00015 (Malignant pleural mesothelioma), H01438 (Neurofibromatosis type 2), H01556 (Meningioma), H01667 (Medulloblastoma)
122786 (FRMD6), 79981 (FRMD1)
60485 (SAV1)
6788 (STK3)
10413 ...
|
HIPPO SIGNALING PATHWAY - MULTIPLE SPECIES
Kibra
NF2
FRMD
SAV1
Mst1/2
YAP/TAZ
Lats1/2
DNA
Mob1
TEAD
YAP/TAZ
Mammals
Kibra
Mer
Salvador
Hippo
Mats
Warts
DNA
Yki
Yki
Drosophila
S. cerevisiae
Cdc15
STE-20
Dbf2
Mob1
Tec1
C ...
|
|
hsa04510
|

|
Focal adhesion
|
Cell-matrix adhesions play essential roles in important biological processes including cell motility, cell proliferation, cell differentiation, regulation of gene expression and cell survival. At the cell-extracellular ...
|
C05981 (Phosphatidylinositol-3,4,5-trisphosphate)
C04637 (1-Phosphatidyl-D-myo-inositol 4,5-bisphosphate)
3688 (ITGB1), 3690 (ITGB3), 3691 (ITGB4), 3693 (ITGB5), 3694 (ITGB6), 3695 (ITGB7), 3696 (ITGB8) ...
|
ECM-receptor interaction
Cytokine-cytokine receptor interaction
MAPK signaling pathway
ITGB
ITGA
Filamin
Cdc42
PIP
PIP
MLCK
MLC
mDia1
MLCP
ROCK
Akt/PKB
PDK1
Parvin
ILK
Vinculin
VASP
PTEN
PI3K
Talin
Zyxin
PIP5K
Actinin
FAK
Calpain
Paxillin
RhoGAP
PKC
Src
Fyn
RhoA
RhoGEF
ECM
Caveolin
FOCAL ...
|
|
hsa04512
|
|
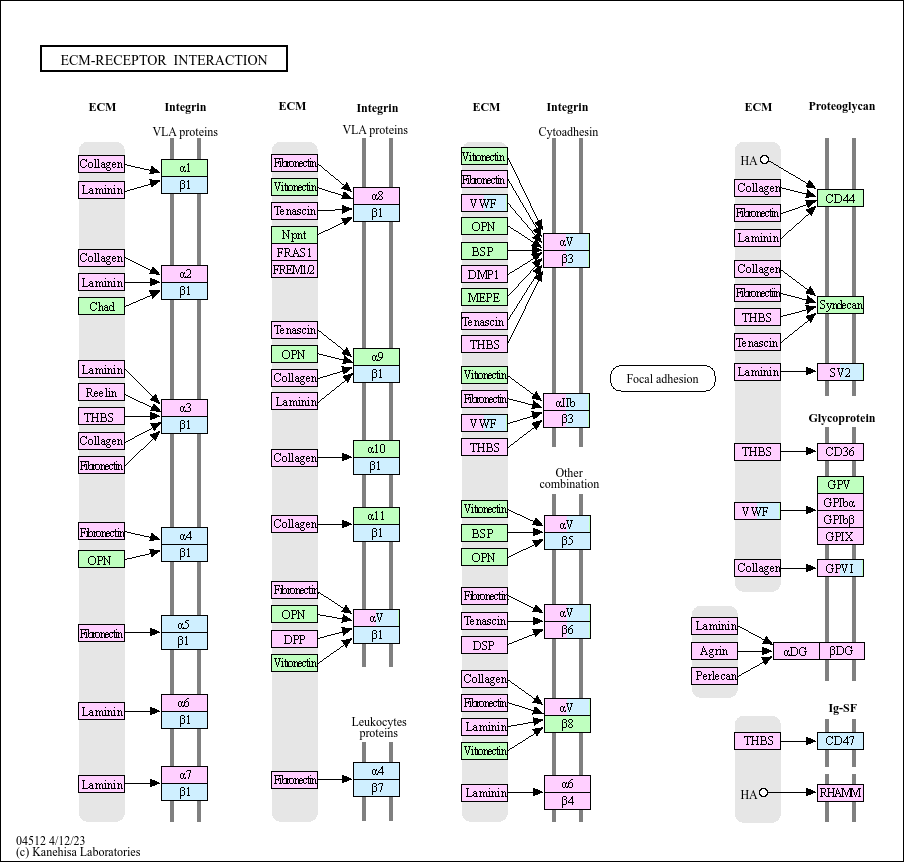
ECM-receptor interaction
|
The extracellular matrix (ECM) consists of a complex mixture of structural and functional macromolecules and serves an important role in tissue and organ morphogenesis and in the maintenance of cell and ...
|
G10505 (Hyaluronic acid)
G10505 (Hyaluronic acid)
10319 (LAMC3), 22798 (LAMB4), 284217 (LAMA1), 3908 (LAMA2), 3909 (LAMA3), 3910 (LAMA4), 3911 (LAMA5), 3912 (LAMB1), 3913 (LAMB2), 3914 (LAMB3), 3915 (LAMC1) ...
|
Laminin
Laminin
Laminin
Laminin
Laminin
Laminin
Laminin
Laminin
Laminin
Laminin
Collagen
Collagen
Collagen
Collagen
Collagen
Collagen
Collagen
Collagen
βDG
Tenascin
GPVI
GPIX
GPIbβ
GPIbα
GPV
VWF
BSP
VWF
Tenascin
Tenascin
Tenascin
Vitronectin
Vitronectin
Vitronectin
Vitronectin
Vitronectin
OPN
OPN
OPN
OPN
Fibronectiin
Fibronectin
Fibronectin
Fibronectin
Fibronectin
Fibronectin
Fibronectin
Fibronectin
Fibronectin
Fibronectin
Fibronectin
THBS
THBS
THBS
THBS
THBS
THBS
Collagen
...
|
|
hsa04514
|

|
Cell adhesion molecules
|
Cell adhesion molecules (CAMs) are (glyco)proteins expressed on the cell surface and play a critical role in a wide array of biologic processes that include hemostasis, the immune response, inflammation ...
|
214 (ALCAM), D11959 (Praluzatamab (USAN/INN))
923 (CD6), D12426 (Itolizumab (USAN/INN))
6693 (SPN)
6614 (SIGLEC1)
5788 (PTPRC), H00091 (T-B+Severe combined immunodeficiency), H00413 (Hepatitis C)
933 (CD22) ...
|
TCR
TCR
TCR
TCR
Leukocyte transendothelial migration
Leukocyte transendothelial migration
T cell receptor signaling pathway
ALCAM
CD6
SPN
PTPRC
CD22
CD8
ITGB1
ITGA8
ITGB2
ITGAL
CD40
CD40
CD40
CD40
NEO1
CDH15
CDH15
CDH4
CDH4
CDH3
CDH3
ITGB1
ITGA6
PVRL2
CDHE
CDHE
CDHE
CDHE
CSPG2
CDHE
CDHE
MAG
MAG
MPZ
MPZ
PVRL3
NFASC
NFASC
CNTNAP2
CNTNAP1
CNTNAP2
CNTNAP1
NF155
NF186
PTPRM
PTPRM
CNTN2
CNTN2
CNTN2
CNTN2
CNTN2
CNTN2
CNTN1
CNTN1
CNTN1
CNTN1
CNTN1
NRCAM
NRCAM
NRCAM
NLGN
β-NRXN
ITGB8
ITGAV
SDC
SDC
PTPRF
NEGR1
NEGR1
IGSF4B
IGSF4B
IGSF4
IGSF4
IGSF4
IGSF4
CDH2
CDH2
CDH2
CDH2
CDH2
CDH2
NCAM
NCAM
L1CAM
L1CAM
L1CAM
L1CAM
NCAM
NCAM
CDH2
CDH2
PVRL1
PVRL1
PVRL3
PVRL3
PVRL1
PVRL1
PVRL3
PVRL3
PVRL1
PVRL1
PVRL2
PVR
CD226
SELE
GLG1
SELP
SELPLG
CD34
GlyCAM1
SELL
MAdCAM1
VCAM1
ITGB7
ITGA9
ITGB1
ITGB1
ITGA4
ITGA4
CD99
CD99
PECAM1
PECAM1
CD58
CD2
ICAM3
ICAM1
ICAM2
JAM2
JAM3
JAM3
ITGB2
ITGAM
ITGB2
ITGAL
JAM1
JAM1
PECAM1
CD40L
JAM1
JAM1
ICAM2
CD40L
SELP
JAM3
ITGB2
CD99
PECAM1
CDH5
ESAM
JAM3
JAM2
JAM1
ITGB2
ITGAL
ICAM1 ...
|
|
hsa04520
|

|
Adherens junction
|
Cell-cell adherens junctions (AJs), the most common type of intercellular adhesions, are important for maintaining tissue architecture and cell polarity and can limit cell movement and proliferation. At ...
|
4089 (SMAD4), H00019 (Pancreatic cancer), H00020 (Colorectal cancer), H00533 (Hereditary hemorrhagic telangiectasia), H01023 (Juvenile polyposis syndrome), H02102 (Myhre syndrome)
999 (CDH1), H00018 (Gastric ...
|
Smad4
E-Cadherin
α-Catenin
p120ctn
p120ctn
E-Cadherin
E-Cadherin
E-Cadherin
TCF/LEF
NLK
TAK1
CBP
Smad3
Yes
Fyn
Fer
Snail
Slug
ERK
TGFβR
FGFR1
ErbB1/2
MET
INSR
IGF-1R
CKII
DEP1
SHP-1
PTP1B
LAR
VEPTP
PTPμ
LMW-PTP
RhoA
RhoA
WAVE
IRSp53
WASP
NWASP
Rac
Rac
Cdc42
Actin
Actin
α-Catenin
ZO-1
ZO-1
Vinculin
Vinculin
Ponsin
α-Actinin
α-Actinin
ADIP
LMO7
Afadin
Afadin
β-Catenin
E-Cadherin
β-Catenin
IQGAP1
IQGAP1
IQGAP1
Rac/cdc42
p120ctn
p120ctn
Src
Nectin
β-Catenin
IQGAP1
Rac/cdc42
Src
Nectin
PAR3
FRG
ADHERENS ...
|
|
hsa04530
|
|
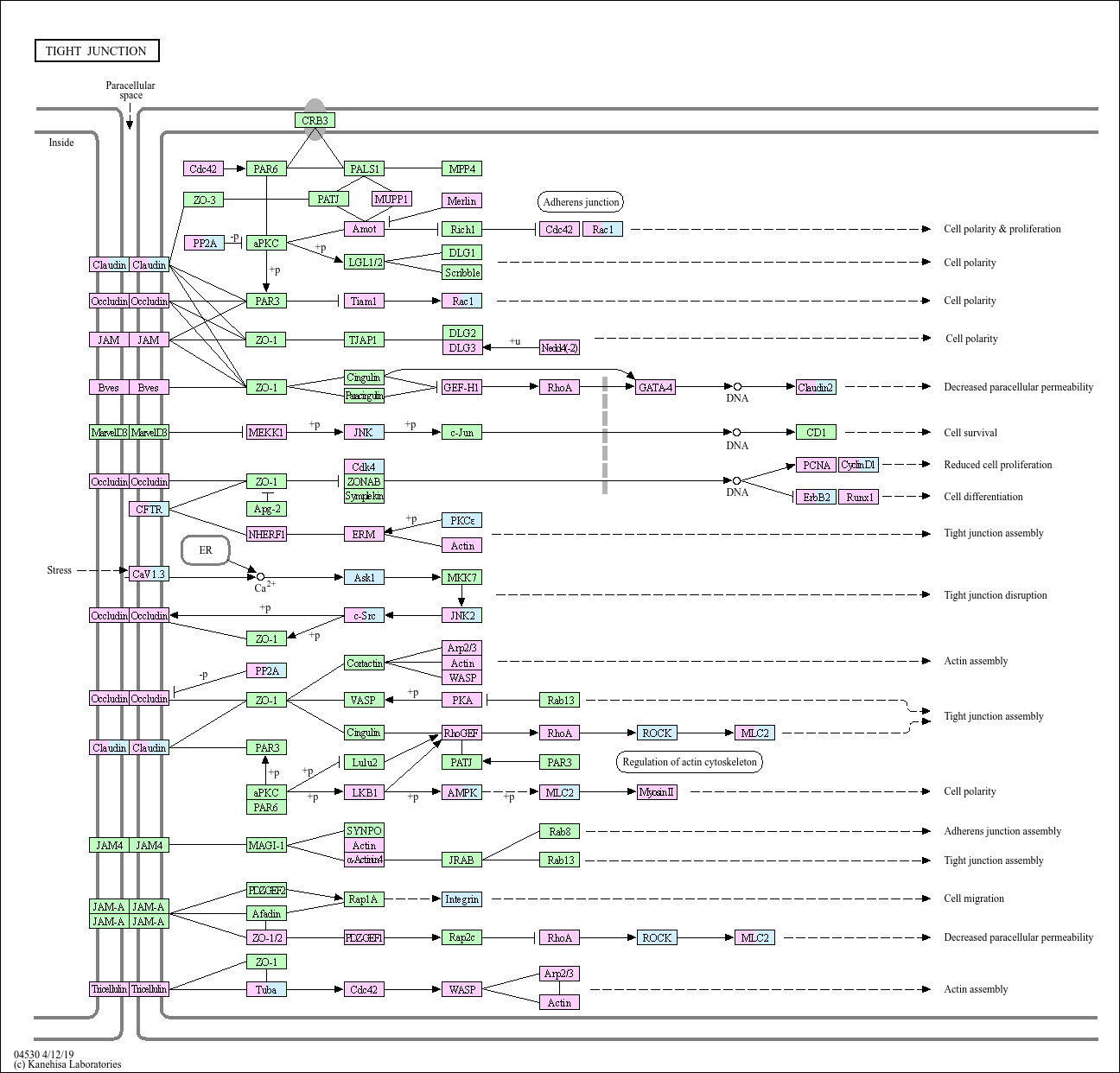
Tight junction
|
Tight junctions (TJs) are essential for establishing a selectively permeable barrier to diffusion through the paracellular space between neighboring cells. TJs are composed of at least three types of transmembrane ...
|
C00076 (Calcium cation)
50855 (PARD6A), 84552 (PARD6G), 84612 (PARD6B)
10207 (PATJ)
998 (CDC42), H02667 (Takenouchi-Kosaki syndrome)
6794 (STK11), H00019 (Pancreatic cancer), H00023 (Testicular cancer) ...
|
PAR6
PATJ
Cdc42
TIGHT JUNCTION
Inside
Paracellular space
LKB1
Cingulin
Tiam1
Decreased paracellular permeability
Rich1
Occludin
ZO-1
Occludin
PATJ
RhoGEF
RhoA
ROCK
MLC2
Lulu2
PAR3
aPKC
Rac1
JAM4
MAGI-1
JAM-A
PDZ-GEF2
Rap1A
Cell ...
|
|
hsa04540
|

|
Gap junction
|
Gap junctions contain intercellular channels that allow direct communication between the cytosolic compartments of adjacent cells. Each gap junction channel is formed by docking of two 'hemichannels', ...
|
C00681 (1-Acyl-sn-glycerol 3-phosphate)
C00076 (Calcium cation)
C01245 (D-myo-Inositol 1,4,5-trisphosphate)
C00942 (3',5'-Cyclic GMP)
C00165 (Diacylglycerol)
C00575 (3',5'-Cyclic AMP)
C00533 (Nitric oxide)
C00780 ...
|
LPA
v-Src
cGMP
DAG
cAMP
CK1
PKC
Serotonin
Glutamate
Connexin
Connexin
Connexin
Connexin
Connexin
Connexin
Connexin
Dopamine
Noradrenaline
ERK5
MEKK2
MEK5
PKG
HTR2
Cdc2
EDG2
PKA
ZO-1
TUBB
ADCY
c-Src
TUBA
Connexin
GUCY
ERK1/2
MEK1/2
Raf-1
Ras
Sos
Grb2
RTK
ADRB1
mGluR
DRD2
PLC
DRD1
IP3R
GAP ...
|
|
hsa04550
|

|
Signaling pathways regulating pluripotency of stem cells
|
Pluripotent stem cells (PSCs) are basic cells with an indefinite self-renewal capacity and the potential to generate all the cell types of the three germinal layers. The types of PSCs known to date include ...
|
51384 (WNT16), 54361 (WNT4), 7471 (WNT1), 7472 (WNT2), 7473 (WNT3), 7474 (WNT5A), 7475 (WNT6), 7476 (WNT7A), 7477 (WNT7B), 7478 (WNT8A), 7479 (WNT8B), 7480 (WNT10B), 7481 (WNT11), 7482 (WNT2B), 7483 (WNT9A) ...
|
Wnt
APC
Axin
Dvl
Frizzled
SIGNALING PATHWAYS REGULATING PLURIPOTENCY OF STEM CELLS
β-catenin
GSK-3β
TCF1
Wnt signaling pathway
DNA
Mouse ES cells (mESCs)
Human ES cells (hESCs)
Mouse EpiS cells (mEpiSCs)
Mouse ...
|
|
hsa04610
|

|
Complement and coagulation cascades
|
The complement system is a proteolytic cascade in blood plasma and a mediator of innate immunity, a nonspecific defense mechanism against pathogens. There are three pathways of complement activation: the ...
|
C00471 (Plasmin)
C03378 (Coagulation factor VIIa)
C03379 (Coagulation factor XIIa)
C03262 (Coagulation factor XIa)
C03259 (Coagulation factor IXa)
C04001 (Coagulation factor VIIIa)
C01065 (Coagulation ...
|
COMPLEMENT AND COAGULATION CASCADES
Plasmin
Coagulation cascade
Extrinsic pathway
Tissue damage
F7a
F12a
F12
F11
F11a
F9a
VWF
F8a
F10a
F10
F2a
(Thrombin)
F5a
Fibrin monomer
Fibrinogen
Fibrin degradation ...
|
|
hsa04611
|

|
Platelet activation
|
Platelets play a key and beneficial role for primary hemostasis on the disruption of the integrity of vessel wall. Platelet adhesion and activation at sites of vascular wall injury is initiated by adhesion ...
|
C00575 (3',5'-Cyclic AMP)
C00076 (Calcium cation)
C01245 (D-myo-Inositol 1,4,5-trisphosphate)
C00165 (Diacylglycerol)
C00076 (Calcium cation)
C00008 (ADP)
C02198 (Thromboxane A2)
C02198 (Thromboxane A2)
C00533 ...
|
PLATELET ACTIVATION
cAMP
DAG
Calcium signaling pathway
Platelet
PAR1/4
P2Y1
ADP
TXA2
PLCβ
GP VI
α2β1
GPV
PLCγ2
RASGRP
Rap1
RIAM
Talin
αIIbβ3
Orai1
Rap1 signaling pathway
PI3K
Akt
eNOS
sGC
PKG
p38
ERK
TXA2
cGMP
G13
Rho-GEF
RhoA
ROCK
MPase
MLC
MLCK
Shape ...
|
|
hsa04612
|

|
Antigen processing and presentation
|
|
972 (CD74), D08944 (Milatuzumab (USAN))
972 (CD74), D08944 (Milatuzumab (USAN))
3108 (HLA-DMA), 3109 (HLA-DMB), 3111 (HLA-DOA), 3112 (HLA-DOB), 3113 (HLA-DPA1), 3115 (HLA-DPB1), 3117 (HLA-DQA1), 3118 (HLA-DQA2) ...
|
CLIP
SLIP
HLA-DM
NFY
RFX
CREB
CIITA
CTSB/L/S
AEP
CTSB
AEP
GILT
TCR
TCR
CD4
KIR
CD8
TAPBP
CALR
ERp57
β2m
β2m
β2M
MHCI
MHCII
MHCII
MHCII
MHCI
MHCI
MHCI
TAP1/2
MHCI
CANX
BiP
HSP90
HSP70
PA28
TNFα
IFNγ
Proteasome
T ...
|
|
hsa04613
|

|
Neutrophil extracellular trap formation
|
Neutrophils play a central role in innate immune defense. One of the mechanisms of neutrophil action is the formation of neutrophil extracellular traps (NETs), the extracellular structures composed of ...
|
C05981 (Phosphatidylinositol-3,4,5-trisphosphate)
C01245 (D-myo-Inositol 1,4,5-trisphosphate)
C00165 (Diacylglycerol)
C00076 (Calcium cation)
C05151 (12-O-Tetradecanoylphorbol 13-acetate)
C00027 (Hydrogen ...
|
Neutrophil
FcγRIIA
FcγRI
Src
Syk
IgG
PI3K
PIP
PLC
DAG
PKC
Raf1
MEK
ERK1/2
Akt
NOD-like receptor signaling pathway
Immune complexes, antibodies, opsonised bacteria
Calcium signaling pathway
NEUTROPHIL ...
|
|
hsa04614
|
|
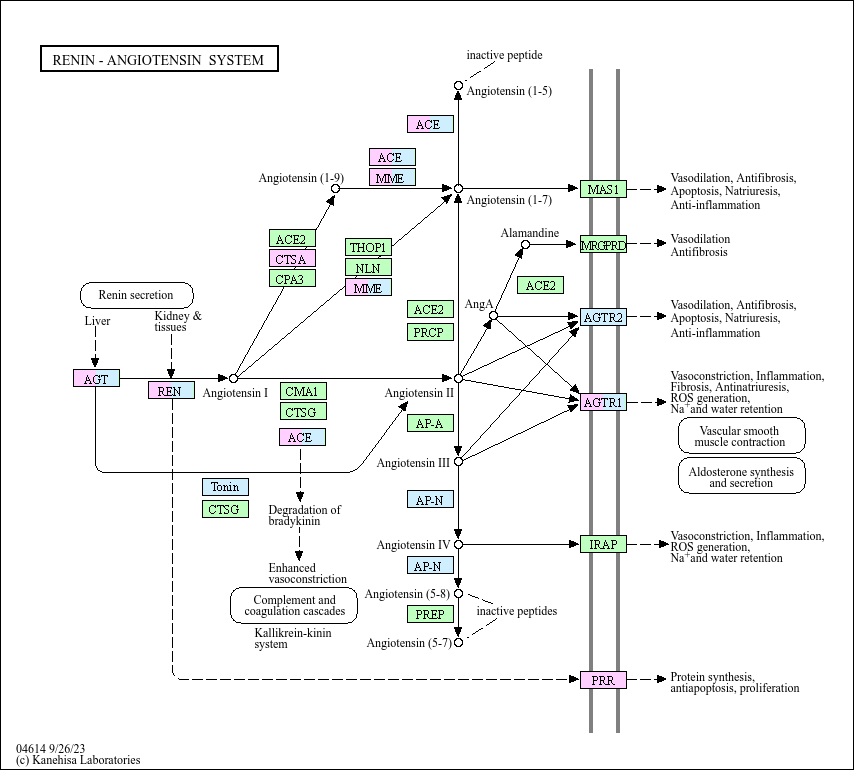
Renin-angiotensin system
|
The renin-angiotensin system (RAS) is a peptidergic system with endocrine characteristics regarding to the regulation of the blood pressure and hydro-electrolytic balance. In the classical RAS, the enzyme ...
|
C02135 (Angiotensin II)
C15849 (Angiotensin IV)
C15848 (Angiotensin III)
C15850 (Angiotensin (1-7))
C15852 (Angiotensin (1-5))
C15851 (Angiotensin (1-9))
C00873 (Angiotensin I)
C20972 (Angiotensin (5-8))
C20973 ...
|
ACE2
ACE2
ACE
ACE
ACE
Angiotensin II
Angiotensin IV
Angiotensin III
Angiotensin (1-7)
Angiotensin (1-5)
Angiotensin (1-9)
Angiotensin I
AGTR1
AGTR2
NLN
THOP1
MME
AP-N
AP-A
CTSA
CPA3
MAS1
MME
IRAP
CMA1
CTSG
AGT
REN
RENIN ...
|
|
hsa04620
|

|
Toll-like receptor signaling pathway
|
Specific families of pattern recognition receptors are responsible for detecting microbial pathogens and generating innate immune responses. Toll-like receptors (TLRs) are membrane-bound receptors identified ...
|
3456 (IFNB1), D12464 (Dazukibart (USAN))
1147 (CHUK), H00882 (Cocoon syndrome), H01931 (Lethal-type popliteal pterygium syndrome)
7189 (TRAF6)
3654 (IRAK1)
51135 (IRAK4), H00096 (Defects of toll-like receptor ...
|
IFN-β
IKKα
TRAF6
IRAK1
IRAK4
MyD88
MyD88
MyD88
TRAF6
ERK
Tpl2
p105
OPN
TRAF3
RIP1
IRF7
IRF5
I-TAC
IP-10
IFN-β
IFN-α
CD86
CD40
IKKβ
TAK1
p38
IRAK1
IKKα
IL-8
TIRAP
MyD88
Rac1
JNK
IL-1β
TBK1
TRAF6
IKKε
FADD
IL-6
RANTES
TAB1
AKT
TAB2
STAT1
IκBα
TNFα
IRF3
CD80
TLR2
MD-2
TLR5
TLR3
TLR2
TLR1
TLR6
LBP
TLR9
TLR4
TRAM
TOLLIP
TRIF
TIRAP
CD14
IKKγ
IRAK4
CASP8
TOLL-LIKE ...
|
|
hsa04621
|

|
NOD-like receptor signaling pathway
|
Specific families of pattern recognition receptors are responsible for detecting various pathogens and generating innate immune responses. The intracellular NOD-like receptor (NLR) family contains more ...
|
C00238 (Potassium cation)
C01245 (D-myo-Inositol 1,4,5-trisphosphate)
C00076 (Calcium cation)
C00002 (ATP)
C00076 (Calcium cation)
C00155 (L-Homocysteine)
C00533 (Nitric oxide)
C00076 (Calcium cation)
29108 ...
|
ASC
pro-CASP1
ASC
pro-CASP1
ASC
pro-CASP1
ASC
pro-CASP1
NOD1
NOD2
iE-DAP
RIP2
IKKα/β
TAK1
TAB
NLRP1
NLRP3
ASC
pro-CASP1
IL-1β
IL-18
Inflammasomes
CARD8
NLRC4
SGT1
HSP90
CASP1
MDP
JNK
ERK
p38
POP2
Pyrin
PSTPIP1
Bacterial ...
|
|
hsa04622
|

|
RIG-I-like receptor signaling pathway
|
Specific families of pattern recognition receptors are responsible for detecting viral pathogens and generating innate immune responses. Non-self RNA appearing in a cell as a result of intracellular viral ...
|
C00046 (RNA)
C00046 (RNA)
23586 (RIGI), H01571 (Singleton-Merten syndrome)
64135 (IFIH1), H00290 (Aicardi-Goutieres syndrome), H00408 (Type 1 diabetes mellitus), H01571 (Singleton-Merten syndrome), H02525 ...
|
RIG-I
MDA5
TRIM25
RNF125
IPS-1
NFκB
DUBA
TRAF3
IRF-3
IKKε
IRF-7
TANK
NAP1
SINTBAD
homo/heterodimer formation
DNA
FADD
CASP8
CASP10
IFNα
IFNβ
TRAF6
IKKα
IKKβ
IKKγ
IκB
Uniquitin-mediated proteolysis
TBK1
LGP2
RNF125
ISG15
Atg5
Atg12
NLRX1
MITA
CYLD
Protein ...
|
|
hsa04623
|

|
Cytosolic DNA-sensing pathway
|
Specific families of pattern recognition receptors are responsible for detecting foreign DNA from invading microbes or host cells and generating innate immune responses. DAI is the first identified sensor ...
|
C20640 (Cyclic GMP-AMP)
C00039 (DNA)
3553 (IL1B), D01583 (Pirfenidone (JAN/USAN/INN)), D06635 (Rilonacept (USAN/INN)), D09315 (Canakinumab (USAN/INN)), D09911 (Gevokizumab (USAN/INN)), D12370 (Lutikizumab ...
|
IL-1β
IL-18
p202
Cell death
ZBP1
TBK1
IKKε/ι
IRF3
Oligomerization
IFNβ
IRF7
IFNα
CXCL10
IPS1
STING
NFκB
RIPK1
RIPK3
ADAR1
E3L
Pyroptosis
Virus
Bacteria
IL-6
Host cells
Shigella flexneri
Listeria monocytogenes
enteropathogenic ...
|
|
hsa04625
|

|
C-type lectin receptor signaling pathway
|
C-type lectin receptors (CLRs) are a large superfamily of proteins characterized by the presence of one or more C-type lectin-like domains (CTLDs). CLRs function as pattern-recognition receptors (PRRs) ...
|
C00965 (1,3-beta-D-Glucan)
C00159 (D-Mannose)
C01019 (L-Fucose)
C00159 (D-Mannose)
C01245 (D-myo-Inositol 1,4,5-trisphosphate)
C00076 (Calcium cation)
C00027 (Hydrogen peroxide), C16844 (Hydroxyl radical)
C21790 ...
|
C-TYPE LECTIN RECEPTOR SIGNALING PATHWAY
Dectin-1
Dectin-2
Mincle
MCL
FcRγ
DC-SIGN
LSP-1
KSR1
CNK1
Src
p50
Raf-1
Pak1
Ras
p65
FcRγ
FcRγ
β-Glucan
Mannose
Syk
IL-23
SHP2
NEMO
IKKα/β
p50
p65
IL-12
NLRP3
Casp1
LSP-1
ASC
IL-1β
pro-IL-1β
Casp8
ASC
IL-6
IL-12
LARG
RHOA
DC-SIGN
LSP-1
Fucose
CYLD
IKKε
STAT1
STAT2
STAT1
IRF9
IL-27
BCL-3
MK2
BCL-3
p50
IL-10
IL-6
IL-12
Syk
SHP2
p50
p65
IL-23
IL-6
pro-IL-1β
Mannose
IκBα
p50
p65
Syk
CARD9
Bcl10
MALT1
p50
p65
IκBα
IL-23
IL-6
IL-12
IL-1β
p50
p65
Ras
PKCδ
IκBα
PLCγ2
ERK
NFAT
IL-23
IL-2
CnA/CnB
IP3R
ROS
NIK
p50
p65
NFAT
p50
p65
TNF
IL-10
Th1 ...
|
|
hsa04630
|

|
JAK-STAT signaling pathway
|
The Janus kinase/signal transducers and activators of transcription (JAK/STAT) pathway is one of a handful of pleiotropic cascades used to transduce a multitude of signals for development and homeostasis ...
|
6772 (STAT1), 6774 (STAT3), 6776 (STAT5A), 6777 (STAT5B), 6778 (STAT6), H00016 (Oral cancer), H00089 (IFN-gamma/IL-12 axis), H00107 (Other well-defined immunodeficiency syndromes), H00931 (Growth hormone ...
|
JAK-STAT SIGNALING PATHWAY
STAT
JAK1/3
Receptor
STAT
IL2 family
STAT
JAK2
Receptor
STAT
Hormone
STAT
JAK2
Receptor
STAT
IL12/23
STAT
JAK1/2
Receptor
STAT
IL6 family
STAT
JAK1
Receptor
STAT
IFN-I/III
Interferons
STAT
JAK1/2
Receptor
STAT
IFN-II
STAT
Receptor
STAT
IL10 ...
|
|
hsa04640
|

|
Hematopoietic cell lineage
|
Blood-cell development progresses from a hematopoietic stem cell (HSC), which can undergo either self-renewal or differentiation into a multilineage committed progenitor cell: a common lymphoid progenitor ...
|
102723407 (IGH), D05994 (Talizumab (USAN/INN))
3655 (ITGA6), 3672 (ITGA1), 3673 (ITGA2), 3675 (ITGA3), 3676 (ITGA4), 3678 (ITGA5), H00586 (Epidermolysis bullosa, junctional), H01235 (Bleeding disorder ...
|
IgM
CD49
CD42
CD41
CD14
CD9
CD61
IL-11R
CD59
CD55
CD35
CD44
CD235a
CD36
CD36
CD117
EPOR
CD125
CD114
CD121
IL-9R
CD14
CD11b
CD11b
CD13
CD13
CD126
CD115
CD64
CD126
CD126
CD124
CD124
CD33
CD116
CD123
CD123
CD123
CD116
CD123
CD116
IgD
CD35
CD23
CD37
CD21
CD20
CD24
CD22
CD19
CD9
CD10
CD3
CD8
CD4
CD1
CD5
CD2
CD7
CD33
CD33
CD33
CD34
CD34
CD34
CD38
CD38
CD71
CD127
CD127
CD25
CD25
CD117
CD117
CD44
TdT
TdT
TdT
CD135
CD34
HLA-DR
HLA-DR
HLA-DR
HLA-DR
HLA-DR
HLA-DR
HLA-DR
TPO
Meg-CSF
TPO
EPO
TNF
M-SCF
G-SCF
G-SCF
Flt3L
Flt3L
Flt3L
Flt3L
Flt3L
Flt3L
IL-11
IL-6
GM-CSF
GM-CSF
GM-CSF
GM-CSF
GM-CSF
GM-CSF
GM-CSF
IL-4
IL-1
IL-11
IL-6
IL-5
IL-4
IL-4
IL-3
IL-3
IL-3
IL-3
IL-3
IL-3
IL-3
IL-3
IL-7
IL-7
SCF
SCF
SCF
SCF
SCF
SCF
SCF
SCF
SCF
SCF
IL-7
HEMATOPOIETIC ...
|







































