| Entry |
|
|---|
| Definition |
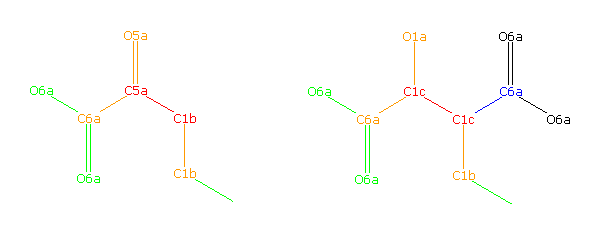
C1b-C1c:*-C6a:C1b+C5a-C1b+C1c
C5a-C1c:*-*:C1b+C6a+O5a-C1c+C6a+O1a
|
|---|
|
 |
|---|
| Reactant pair |
|
|---|
| Related class |
|
|---|
| Reaction |
|
|---|
| Enzyme |
|
|---|
| Pathway |
| rn00400 | Phenylalanine, tyrosine and tryptophan biosynthesis |
| rn00440 | Phosphonate and phosphinate metabolism |
| rn00720 | Other carbon fixation pathways |
| rn01110 | Biosynthesis of secondary metabolites |
| rn01120 | Microbial metabolism in diverse environments |
| rn01210 | 2-Oxocarboxylic acid metabolism |
| rn01230 | Biosynthesis of amino acids |
|
|---|
| RModule |
| RM001 | 2-Oxocarboxylic acid chain extension by tricarboxylic acid pathway |
|
|---|
| Orthology |
| K17750 | 3-benzylmalate dehydrogenase [EC:1.1.1.-] |
| K21360 | 3-isopropylmalate/methylthioalkylmalate dehydrogenase [EC:1.1.1.85 1.1.1.-] |
|
|---|