| Entry |
|
|---|
| Name |
Riboflavin metabolism - Klebsiella pneumoniae Kp52.145 |
|---|
| Class |
Metabolism; Metabolism of cofactors and vitamins
|
|---|
| Pathway map |
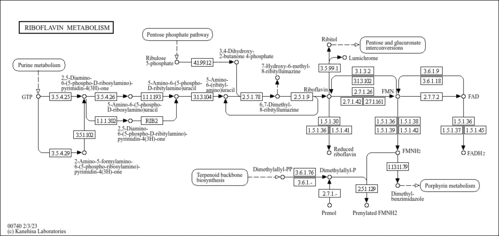
|
|---|
| Module |
| kpnk_M00125 | Riboflavin biosynthesis, plants and bacteria, GTP => riboflavin/FMN/FAD [PATH:kpnk00740] |
|
|---|
| Other DBs |
|
|---|
| Organism |
Klebsiella pneumoniae Kp52.145 [GN: kpnk]
|
|---|
| Gene |
| BN49_0654 | ribB; highly similar to 3,4-dihydroxy-2-butanone 4-phosphate synthase from Klebsiella pneumoniae subsp. pneumoniae strain ATCC 700721 - MGH 78578 (sp|A6TE29) [KO:K02858] [EC:4.1.99.12] |
| BN49_0660 | yqiE; highly similar to ADP-ribose pyrophosphatase from Escherichia coli strain UTI89 - UPEC (tr|Q1R6U9) [KO:K01515] [EC:3.6.1.13 3.6.1.-] |
| BN49_1028 | pad1; highly similar to probable aromatic acid decarboxylase from Escherichia coli O157:H7 (sp|P69772) [KO:K03186] [EC:2.5.1.129] |
| BN49_1340 | ribD; highly similar to riboflavin biosynthesis protein from Escherichia coli O44:H18 strain 042 - EAEC (tr|D3GUA9) [KO:K11752] [EC:3.5.4.26 1.1.1.193] |
| BN49_1341 | ribH; highly similar to 6,7-dimethyl-8-ribityllumazine synthase from Shigella sonnei (sp|Q3Z4Z4) [KO:K00794] [EC:2.5.1.78] |
| BN49_2068 | ssuE; highly similar to FMN reductase from Escherichia coli strain K12 (sp|P80644) [KO:K00299] [EC:1.5.1.38] |
| BN49_2340 | phoN; highly similar to non-specific acid phosphatase from Providencia stuartii (sp|P26975) [KO:K09474] [EC:3.1.3.2] |
| BN49_2412 | ribA; highly similar to GTP cyclohydrolase-2 from Klebsiella pneumoniae subsp. pneumoniae strain ATCC 700721 - MGH 78578 (sp|A6T7Y5) [KO:K01497] [EC:3.5.4.25] |
| BN49_3097 | ribE; highly similar to riboflavin synthase alpha chain from Escherichia coli O157:H7 (tr|Q8X622) [KO:K00793] [EC:2.5.1.9] |
| BN49_3768 | thiM; highly similar to hydroxyethylthiazole kinase from Klebsiella pneumoniae strain 342 (sp|B5XP99) [KO:K00878] [EC:2.7.1.50] |
| BN49_3918 | ubiX; highly similar to 3-octaprenyl-4-hydroxybenzoate carboxy-lyase from Escherichia coli strain K12 (sp|P0AG03) [KO:K03186] [EC:2.5.1.129] |
| BN49_4321 | ribF; highly similar to bifunctional riboflavin kinase-FAD synthetase from Escherichia coli O127:H6 strain E2348-69 - EPEC (tr|B7UI70) [KO:K11753] [EC:2.7.1.26 2.7.7.2] |
| BN49_4415 | hpaC; highly similar to 4-hydroxyphenylacetate 3-monooxygenase reductase component from Escherichia coli (sp|Q57501) [KO:K00484] [EC:1.5.1.36] |
| BN49_4501 | fre; highly similar to NAD(P)H-flavin reductase from Salmonella typhimurium (sp|Q9L6L9) [KO:K05368] [EC:1.5.1.41] |
| BN49_4990 | aphA; highly similar to diadenosine tetraphosphatase from Escherichia coli O1:K1 - APEC (tr|A1AIP5) [KO:K03788] [EC:3.1.3.2] |
|
|---|
| Compound |
| C00235 | Dimethylallyl diphosphate |
| C01268 | 5-Amino-6-(5'-phosphoribosylamino)uracil |
| C01304 | 2,5-Diamino-6-(5-phospho-D-ribosylamino)pyrimidin-4(3H)-one |
| C04332 | 6,7-Dimethyl-8-(D-ribityl)lumazine |
| C04454 | 5-Amino-6-(5'-phospho-D-ribitylamino)uracil |
| C04732 | 5-Amino-6-(1-D-ribitylamino)uracil |
| C05995 | 7-Hydroxy-6-methyl-8-ribityllumazine |
| C15556 | L-3,4-Dihydroxybutan-2-one 4-phosphate |
| C15563 | 2-Amino-5-formylamino-6-(5-phospho-D-ribosylamino)pyrimidin-4(3H)-one |
| C18910 | 2,5-Diamino-6-(5-phospho-D-ribitylamino)pyrimidin-4(3H)-one |
| C21214 | Dimethylallyl phosphate |
|
|---|
| Reference |
|
|---|
| Authors |
Vitreschak AG, Rodionov DA, Mironov AA, Gelfand MS. |
|---|
| Title |
Regulation of riboflavin biosynthesis and transport genes in bacteria by transcriptional and translational attenuation. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Fischer M, Haase I, Feicht R, Richter G, Gerhardt S, Changeux JP, Huber R, Bacher A. |
|---|
| Title |
Biosynthesis of riboflavin: 6,7-dimethyl-8-ribityllumazine synthase of Schizosaccharomyces pombe. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Haase I, Fischer M, Bacher A, Schramek N. |
|---|
| Title |
Temperature-dependent presteady state kinetics of lumazine synthase from the hyperthermophilic eubacterium Aquifex aeolicus. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Fischer M, Romisch W, Illarionov B, Eisenreich W, Bacher A. |
|---|
| Title |
Structures and reaction mechanisms of riboflavin synthases of eubacterial and archaeal origin. |
|---|
| Journal |
|
|---|
Related
pathway |
| kpnk00040 | Pentose and glucuronate interconversions |
|
|---|
| KO pathway |
|
|---|