| Entry |
|
|---|
| Name |
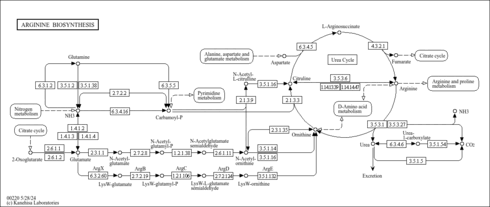
Arginine biosynthesis - Photorhabdus laumondii subsp. laumondii TTO1 |
|---|
| Class |
Metabolism; Amino acid metabolism
|
|---|
| Pathway map |
|
|---|
| Module |
|
|---|
| Other DBs |
|
|---|
| Organism |
Photorhabdus laumondii subsp. laumondii TTO1 [GN: plu]
|
|---|
| Gene |
| plu1172 | unnamed protein product; Highly similar to probable glutaminase YneH of Escherichia coli [KO:K01425] [EC:3.5.1.2] |
| plu4743 | argB; acetylglutamate kinase (NAG kinase) (AGK) (N-acetyl-L-glutamate 5-phosphotransferase) [KO:K00930] [EC:2.7.2.8] |
| plu4744 | argC; N-acetyl-gamma-glutamyl-phosphate reductase (AGPR) (N-acetyl-glutamate semialdehyde dehydrogenase) (NAGSA dehydrogenase) [KO:K00145] [EC:1.2.1.38] |
| plu4745 | argE; acetylornithine deacetylase (acetylornithinase) (AO) (N-acetylornithinase) (NAO) [KO:K01438] [EC:3.5.1.16] |
|
|---|
| Compound |
| C01250 | N-Acetyl-L-glutamate 5-semialdehyde |
| C03406 | N-(L-Arginino)succinate |
| C04133 | N-Acetyl-L-glutamate 5-phosphate |
| C20949 | LysW-L-glutamyl 5-phosphate |
| C20950 | LysW-L-glutamate 5-semialdehyde |
|
|---|
Related
pathway |
| plu00250 | Alanine, aspartate and glutamate metabolism |
| plu00330 | Arginine and proline metabolism |
|
|---|
| KO pathway |
|
|---|
| LinkDB |
|
|---|