| Entry |
|
|---|
| Name |
enzyme N6-(dihydrolipoyl)lysine:NAD+ oxidoreductase
|
|---|
| Definition |
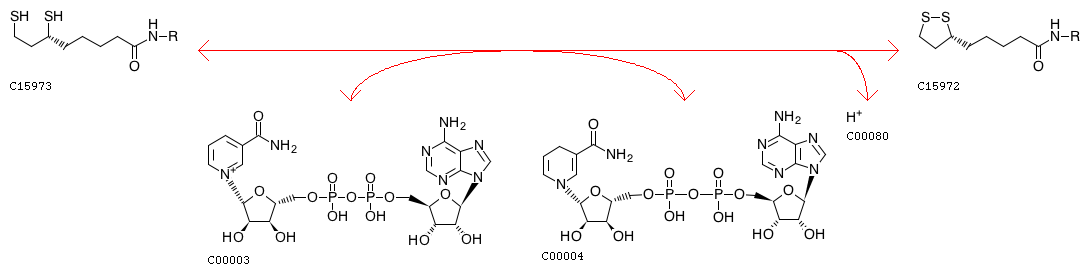
Enzyme N6-(dihydrolipoyl)lysine + NAD+ <=> Enzyme N6-(lipoyl)lysine + NADH + H+
|
|---|
| Equation |
|
|---|
|
 |
|---|
| Comment |
Oxo-acid dehydrogenase complexes, dihydrolipoyl dehydrogenase
|
|---|
| Reaction class |
|
|---|
| Enzyme |
|
|---|
| Pathway |
| rn00010 | Glycolysis / Gluconeogenesis |
| rn00280 | Valine, leucine and isoleucine degradation |
| rn01110 | Biosynthesis of secondary metabolites |
|
|---|
| Module |
| M00009 | Citrate cycle (TCA cycle, Krebs cycle) |
| M00011 | Citrate cycle, second carbon oxidation, 2-oxoglutarate => oxaloacetate |
| M00032 | Lysine degradation, lysine => saccharopine => acetoacetyl-CoA |
| M00036 | Leucine degradation, leucine => acetoacetate + acetyl-CoA |
| M00307 | Pyruvate oxidation, pyruvate => acetyl-CoA |
|
|---|
| Brite |
Enzymatic reactions [BR:br08201]
1. Oxidoreductase reactions
1.8 Acting on a sulfur group of donors
1.8.1 With NAD+ or NADP+ as acceptor
1.8.1.4
R07618 Enzyme N6-(dihydrolipoyl)lysine + NAD+ <=> Enzyme N6-(lipoyl)lysine + NADH + H+
IUBMB reaction hierarchy [BR:br08202]
1. Oxidoreductase reactions
1.8.1.4
R08550 [Protein]-N6-[(R)-dihydrolipoyl]-L-lysine <=> Protein N6-(lipoyl)lysine
R07618 Enzyme N6-(dihydrolipoyl)lysine <=> Enzyme N6-(lipoyl)lysine
|
|---|
| Orthology |
|
|---|
| Other DBs |
|
|---|
| LinkDB |
|
|---|