| Entry |
|
|---|
| Name |
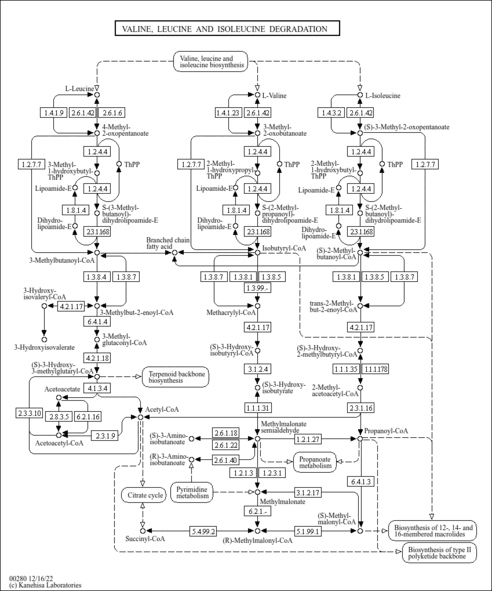
Valine, leucine and isoleucine degradation - Bacillus velezensis SQR9 |
|---|
| Class |
Metabolism; Amino acid metabolism
|
|---|
| Pathway map |
| bamy00280 | Valine, leucine and isoleucine degradation |
|
|---|
| Other DBs |
|
|---|
| Organism |
Bacillus velezensis SQR9 [GN: bamy]
|
|---|
| Gene |
|
|---|
| Compound |
| C00141 | 3-Methyl-2-oxobutanoic acid |
| C00233 | 4-Methyl-2-oxopentanoate |
| C00349 | 2-Methyl-3-oxopropanoate |
| C00356 | (S)-3-Hydroxy-3-methylglutaryl-CoA |
| C00671 | (S)-3-Methyl-2-oxopentanoic acid |
| C01205 | (R)-3-Amino-2-methylpropanoate |
| C03344 | 2-Methylacetoacetyl-CoA |
| C03345 | 2-Methylbut-2-enoyl-CoA |
| C03460 | 2-Methylprop-2-enoyl-CoA |
| C04405 | (2S,3S)-3-Hydroxy-2-methylbutanoyl-CoA |
| C05996 | Branched chain fatty acid |
| C05998 | 3-Hydroxyisovaleryl-CoA |
| C06000 | (S)-3-Hydroxyisobutyryl-CoA |
| C06001 | (S)-3-Hydroxyisobutyrate |
| C06002 | (S)-Methylmalonate semialdehyde |
| C15972 | Enzyme N6-(lipoyl)lysine |
| C15973 | Enzyme N6-(dihydrolipoyl)lysine |
| C15974 | 3-Methyl-1-hydroxybutyl-ThPP |
| C15975 | [Dihydrolipoyllysine-residue (2-methylpropanoyl)transferase] S-(3-methylbutanoyl)dihydrolipoyllysine |
| C15976 | 2-Methyl-1-hydroxypropyl-ThPP |
| C15977 | [Dihydrolipoyllysine-residue (2-methylpropanoyl)transferase] S-(2-methylpropanoyl)dihydrolipoyllysine |
| C15978 | 2-Methyl-1-hydroxybutyl-ThPP |
| C15979 | [Dihydrolipoyllysine-residue (2-methylpropanoyl)transferase] S-(2-methylbutanoyl)dihydrolipoyllysine |
| C15980 | (S)-2-Methylbutanoyl-CoA |
|
|---|
Related
pathway |
| bamy00290 | Valine, leucine and isoleucine biosynthesis |
|
|---|
| KO pathway |
|
|---|