| Entry |
|
|---|
| Name |
Phenylalanine, tyrosine and tryptophan biosynthesis - Polymorphobacter sp. PAMC 29334 |
|---|
| Class |
Metabolism; Amino acid metabolism
|
|---|
| Pathway map |
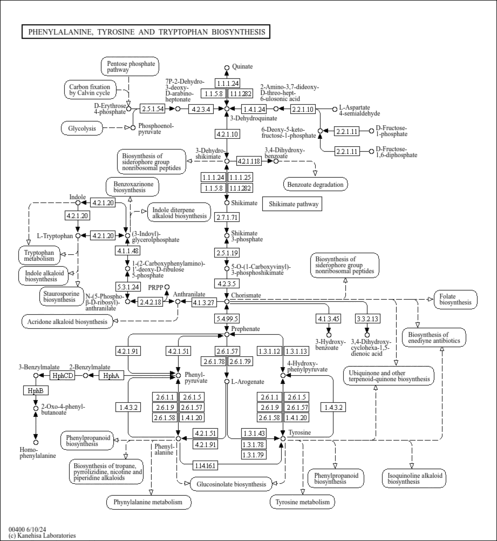
| poq00400 | Phenylalanine, tyrosine and tryptophan biosynthesis |
|
|---|
| Module |
| poq_M00022 | Shikimate pathway, phosphoenolpyruvate + erythrose-4P => chorismate [PATH:poq00400] |
|
|---|
| Other DBs |
|
|---|
| Organism |
Polymorphobacter sp. PAMC 29334 [GN: poq]
|
|---|
| Gene |
|
|---|
| Compound |
| C00119 | 5-Phospho-alpha-D-ribose 1-diphosphate |
| C00279 | D-Erythrose 4-phosphate |
| C00354 | D-Fructose 1,6-bisphosphate |
| C00441 | L-Aspartate 4-semialdehyde |
| C01179 | 3-(4-Hydroxyphenyl)pyruvate |
| C01269 | 5-O-(1-Carboxyvinyl)-3-phosphoshikimate |
| C01302 | 1-(2-Carboxyphenylamino)-1-deoxy-D-ribulose 5-phosphate |
| C03506 | Indoleglycerol phosphate |
| C04302 | N-(5-Phospho-D-ribosyl)anthranilate |
| C04691 | 2-Dehydro-3-deoxy-D-arabino-heptonate 7-phosphate |
| C16848 | 6-Deoxy-5-ketofructose 1-phosphate |
| C16850 | 2-Amino-3,7-dideoxy-D-threo-hept-6-ulosonic acid |
| C20327 | 2-Oxo-4-phenylbutyric acid |
| C20710 | (4R,5R)-4,5-Dihydroxycyclohexa-1(6),2-diene-1-carboxylate |
|
|---|
Related
pathway |
| poq00130 | Ubiquinone and other terpenoid-quinone biosynthesis |
| poq00710 | Carbon fixation by Calvin cycle |
|
|---|
| KO pathway |
|
|---|