| Entry |
|
|---|
| Name |
Pantothenate and CoA biosynthesis - Anser cygnoides (swan goose) |
|---|
| Class |
Metabolism; Metabolism of cofactors and vitamins
|
|---|
| Pathway map |
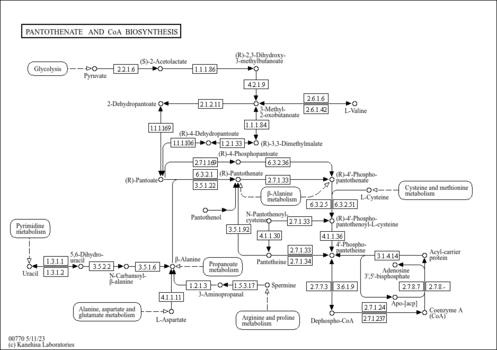
|
|---|
| Module |
| acyg_M00046 | Pyrimidine degradation, uracil => beta-alanine, thymine => 3-aminoisobutanoate [PATH:acyg00770] |
|
|---|
| Other DBs |
|
|---|
| Organism |
Anser cygnoides (swan goose) [GN: acyg]
|
|---|
| Gene |
| 106047540 | AASDHPPT; L-aminoadipate-semialdehyde dehydrogenase-phosphopantetheinyl transferase isoform X1 [KO:K06133] [EC:2.7.8.-] |
|
|---|
| Compound |
| C00054 | Adenosine 3',5'-bisphosphate |
| C00141 | 3-Methyl-2-oxobutanoic acid |
| C01134 | Pantetheine 4'-phosphate |
| C03492 | D-4'-Phosphopantothenate |
| C03688 | Apo-[acyl-carrier protein] |
| C04079 | N-((R)-Pantothenoyl)-L-cysteine |
| C04272 | (R)-2,3-Dihydroxy-3-methylbutanoate |
| C04352 | (R)-4'-Phosphopantothenoyl-L-cysteine |
|
|---|
Related
pathway |
| acyg00250 | Alanine, aspartate and glutamate metabolism |
|
|---|
| KO pathway |
|
|---|