| Entry |
|
|---|
| Name |
Arginine biosynthesis - Kytococcus sedentarius |
|---|
| Class |
Metabolism; Amino acid metabolism
|
|---|
| Pathway map |
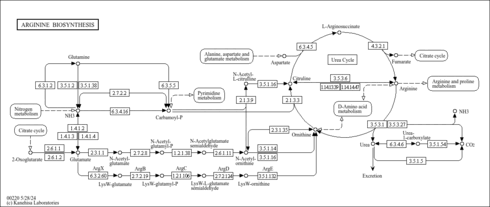
|
|---|
| Module |
|
|---|
| Other DBs |
|
|---|
| Organism |
Kytococcus sedentarius [GN: kse]
|
|---|
| Gene |
|
|---|
| Compound |
| C01250 | N-Acetyl-L-glutamate 5-semialdehyde |
| C03406 | N-(L-Arginino)succinate |
| C04133 | N-Acetyl-L-glutamate 5-phosphate |
| C20949 | LysW-L-glutamyl 5-phosphate |
| C20950 | LysW-L-glutamate 5-semialdehyde |
|
|---|
Related
pathway |
| kse00250 | Alanine, aspartate and glutamate metabolism |
| kse00330 | Arginine and proline metabolism |
|
|---|
| KO pathway |
|
|---|