 | | PATHWAY: bmf00240 | |
| Entry |
|
|---|
| Name |
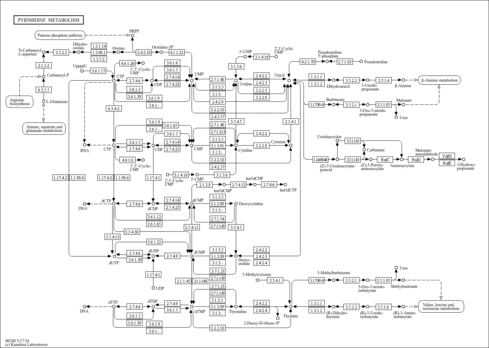
Pyrimidine metabolism - Brucella abortus 2308 |
|---|
| Class |
Metabolism; Nucleotide metabolism
|
|---|
| Pathway map |
|
|---|
| Module |
| bmf_M00053 | Deoxyribonucleotide biosynthesis, ADP/GDP/CDP/UDP => dATP/dGTP/dCTP/dUTP [PATH:bmf00240] |
|
|---|
| Other DBs |
|
|---|
| Organism |
Brucella abortus 2308 [GN: bmf]
|
|---|
| Gene |
| BAB1_1508 | Methylglyoxal synthase-like domain:Carbamoyl-phosphate synthase L chain, ATP-binding:Carbamoyl-phosphate synthetase large cha... [KO:K01955] [EC:6.3.5.5] |
| BAB1_1502 | Glutamine amidotransferase class-I:Carbamoyl-phosphate synthase, GATase domain:Carbamoyl-phosphate synthase, small chain:Anth... [KO:K01956] [EC:6.3.5.5] |
| BAB2_0641 | pyrB; Aspartate carbamoyltransferase:Ornithine carbamoyltransferase:Aspartate/ornithine carbamoyltransferase:Aspartate/ornithine ca... [KO:K00609] [EC:2.1.3.2] |
| BAB1_0688 | pyrC-1; Dihydroorotase:Dihydroorotase multifunctional complex type:Amidohydrolase [KO:K01465] [EC:3.5.2.3] |
| BAB1_0341 | pyrD; Dihydroorotate dehydrogenase:FMN/related compound-binding core:Dihydroorotate dehydrogenase 2 [KO:K00254] [EC:1.3.5.2] |
| BAB1_0673 | pyrE; Phosphoribosyltransferase:Purine/pyrimidine phosphoribosyl transferase:Orotate phosphoribosyltransferase, Thermus type [KO:K00762] [EC:2.4.2.10] |
| BAB2_0889 | nrdE; Ribonucleotide reductase large subunit:Prenyl group binding site (CAAX box) [KO:K00525] [EC:1.17.4.1] |
| BAB1_1687 | dut; DeoxyUTP pyrophosphatase:DeoxyUTP pyrophosphatase, subfamily 1:DeoxyUTP pyrophosphatase, subfamily 2 [KO:K01520] [EC:3.6.1.23] |
| BAB1_0314 | Pyridine nucleotide-disulphide oxidoreductase, class-II:NAD binding site:Adrenodoxin reductase:Pyridine nucleotide-disulphide... [KO:K17722] [EC:1.3.1.1] |
| BAB1_0313 | Legume lectin, beta domain:Dihydroorotate dehydrogenase:4Fe-4S ferredoxin, iron-sulfur binding domain:FMN/related compound-bi... [KO:K17723] [EC:1.3.1.1] |
|
|---|
| Compound |
| C00119 | 5-Phospho-alpha-D-ribose 1-diphosphate |
| C00438 | N-Carbamoyl-L-aspartate |
| C00672 | 2-Deoxy-D-ribose 1-phosphate |
| C01168 | Pseudouridine 5'-phosphate |
| C01205 | (R)-3-Amino-2-methylpropanoate |
| C03997 | 5-Hydroxymethyldeoxycytidylate |
| C06198 | P1,P4-Bis(5'-uridyl) tetraphosphate |
| C11038 | 2'-Deoxy-5-hydroxymethylcytidine-5'-diphosphate |
| C11039 | 2'-Deoxy-5-hydroxymethylcytidine-5'-triphosphate |
| C15607 | 3-Oxo-3-ureidopropanoate |
| C20231 | (Z)-3-Ureidoacrylate peracid |
| C20249 | (Z)-3-Peroxyaminoacrylate |
| C20846 | 3-Oxo-3-ureidoisobutyrate |
| C21029 | (R)-3-Ureidoisobutyrate |
|
|---|
Related
pathway |
| bmf00250 | Alanine, aspartate and glutamate metabolism |
| bmf00280 | Valine, leucine and isoleucine degradation |
|
|---|
| KO pathway |
|
|---|
| LinkDB |
|
|---|
|
|