 | | PATHWAY: bmf00650 | |
| Entry |
|
|---|
| Name |
Butanoate metabolism - Brucella abortus 2308 |
|---|
| Class |
Metabolism; Carbohydrate metabolism
|
|---|
| Pathway map |
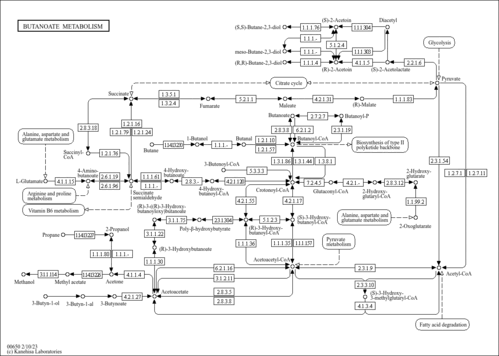
|
|---|
| Module |
|
|---|
| Other DBs |
|
|---|
| Organism |
Brucella abortus 2308 [GN: bmf]
|
|---|
| Gene |
| BAB2_1045 | NAD binding site:FAD-dependent pyridine nucleotide-disulphide oxidoreductase:3-hydroxyacyl-CoA dehydrogenase, C-terminal:3-hy... [KO:K00074] [EC:1.1.1.157] |
| BAB1_1970 | 3-hydroxyacyl-CoA dehydrogenase, C-terminal:3-hydroxyacyl-CoA dehydrogenase, NAD binding domain:3-hydroxyacyl-CoA dehydrogenase [KO:K00074] [EC:1.1.1.157] |
| BAB2_0791 | Acyl-CoA dehydrogenase:Acyl-CoA dehydrogenase, C-terminal:Acyl-CoA dehydrogenase, central domain:Acyl-CoA dehydrogenase, N-te... [KO:K00248] [EC:1.3.8.1] |
| BAB1_1901 | FAD-dependent pyridine nucleotide-disulphide oxidoreductase:Fumarate reductase/succinate dehydrogenase, FAD-binding site:Fuma... [KO:K00239] [EC:1.3.5.1] |
| BAB1_1900 | 4Fe-4S ferredoxin, iron-sulfur binding domain:Succinate dehydrogenase/fumarate reductase iron-sulfur protein:2Fe-2S ferredoxin [KO:K00240] [EC:1.3.5.1] |
| BAB2_0865 | Bacterial extracellular solute-binding protein, family 3:Pyridoxal-dependent decarboxylase [KO:K01580] [EC:4.1.1.15] |
| BAB1_1789 | Eukaryotic thiol (cysteine) protease:Short-chain dehydrogenase/reductase SDR:Glucose/ribitol dehydrogenase [KO:K00019] [EC:1.1.1.30] |
| BAB1_1408 | ilvB; Pyruvate decarboxylase:Acetolactate synthase large subunit biosynthetic type [KO:K01652] [EC:2.2.1.6] |
|
|---|
| Compound |
| C00356 | (S)-3-Hydroxy-3-methylglutaryl-CoA |
| C01144 | (S)-3-Hydroxybutanoyl-CoA |
| C03561 | (R)-3-Hydroxybutanoyl-CoA |
| C04546 | (R)-3-((R)-3-Hydroxybutanoyloxy)butanoate |
| C06143 | Poly-beta-hydroxybutyrate |
|
|---|
Related
pathway |
| bmf00250 | Alanine, aspartate and glutamate metabolism |
| bmf00330 | Arginine and proline metabolism |
|
|---|
| KO pathway |
|
|---|
| LinkDB |
|
|---|
|
|