 | | PATHWAY: cdet00640 | |
| Entry |
|
|---|
| Name |
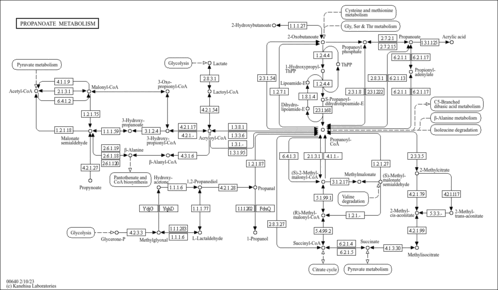
Propanoate metabolism - Colletotrichum destructivum |
|---|
| Class |
Metabolism; Carbohydrate metabolism
|
|---|
| Pathway map |
|
|---|
| Module |
|
|---|
| Other DBs |
|
|---|
| Organism |
Colletotrichum destructivum [GN: cdet]
|
|---|
| Gene |
| 87936933 | CDEST_00430; Putative enoyl-CoA hydratase/isomerase, ClpP/crotonase-like domain superfamily [KO:K07511] [EC:4.2.1.17] |
| 87937711 | CDEST_01208; Putative enoyl-CoA hydratase/isomerase, ClpP/crotonase-like domain superfamily [KO:K07511] [EC:4.2.1.17] |
| 87939243 | CDEST_02740; Putative biotin/lipoyl attachment, 2-oxoacid dehydrogenase acyltransferase, catalytic [KO:K09699] [EC:2.3.1.168] |
| 87939878 | CDEST_03375; Putative pyridine nucleotide-disulfide oxidoreductase, class I, FAD/NAD(P)-binding protein [KO:K00382] [EC:1.8.1.4] |
| 87941211 | CDEST_04708; Putative enoyl-CoA hydratase/isomerase, ClpP/crotonase-like domain superfamily [KO:K05605] [EC:3.1.2.4] |
| 87943319 | CDEST_06816; Putative succinate--CoA synthetase, beta subunit, ATP-citrate lyase/succinyl-CoA ligase [KO:K01900] [EC:6.2.1.4 6.2.1.5] |
| 87943333 | CDEST_06830; Putative AMP-dependent synthetase/ligase domain, acetate-CoA ligase, AMP-binding, ANL [KO:K01895] [EC:6.2.1.1] |
| 87944922 | CDEST_08419; Putative short-chain dehydrogenase/reductase SDR, MaoC-like dehydratase domain-containing protein [KO:K14729] [EC:4.2.1.- 1.1.1.-] |
| 87945270 | CDEST_08767; Putative transketolase-like, pyrimidine-binding domain, thiamine diphosphate-binding protein [KO:K00167] [EC:1.2.4.4] |
| 87945304 | CDEST_08801; Putative dehydrogenase, E1 component, thiamine diphosphate-binding protein [KO:K00166] [EC:1.2.4.4] |
| 87946019 | CDEST_09516; Putative acyl-CoA oxidase, acyl-CoA dehydrogenase/oxidase and middle domain superfamily [KO:K00232] [EC:1.3.3.6] |
| 87946603 | CDEST_10101; Putative aliphatic acid kinase, short-chain, ATPase, nucleotide binding domain-containing protein [KO:K00925] [EC:2.7.2.1] |
| 87947076 | CDEST_10576; Putative 6-phosphogluconate dehydrogenase, NADP-binding, 6-phosphogluconate dehydrogenase, domain 2 [KO:K23146] |
| 87949939 | CDEST_13439; Putative succinate--CoA synthetase, beta subunit, ATP-citrate lyase/succinyl-CoA ligase [KO:K01900] [EC:6.2.1.4 6.2.1.5] |
| 87949978 | CDEST_13478; Putative coA-binding, succinyl-CoA ligase, alpha subunit, ATP-citrate lyase/succinyl-CoA ligase [KO:K01899] [EC:6.2.1.4 6.2.1.5] |
| 87950201 | CDEST_13701; Putative AMP-dependent synthetase/ligase domain, AMP-binding, AMP-binding enzyme domain, ANL [KO:K01908] [EC:6.2.1.17] |
| 87951524 | CDEST_15024; Putative coA-binding, succinyl-CoA ligase, alpha subunit, ATP-citrate lyase/succinyl-CoA ligase [KO:K01899] [EC:6.2.1.4 6.2.1.5] |
|
|---|
| Compound |
| C04225 | (Z)-But-2-ene-1,2,3-tricarboxylate |
| C04593 | (2S,3R)-3-Hydroxybutane-1,2,3-tricarboxylate |
| C06002 | (S)-Methylmalonate semialdehyde |
| C15972 | Enzyme N6-(lipoyl)lysine |
| C15973 | Enzyme N6-(dihydrolipoyl)lysine |
| C21017 | 2-(alpha-Hydroxypropyl)thiamine diphosphate |
| C21018 | Enzyme N6-(S-propyldihydrolipoyl)lysine |
| C21250 | 2-Methyl-trans-aconitate |
|
|---|
Related
pathway |
| cdet00260 | Glycine, serine and threonine metabolism |
| cdet00280 | Valine, leucine and isoleucine degradation |
| cdet00660 | C5-Branched dibasic acid metabolism |
|
|---|
| KO pathway |
|
|---|
|
|