| Entry |
|
|---|
| Name |
Pyrimidine metabolism - Azospirillum ramasamyi |
|---|
| Class |
Metabolism; Nucleotide metabolism
|
|---|
| Pathway map |
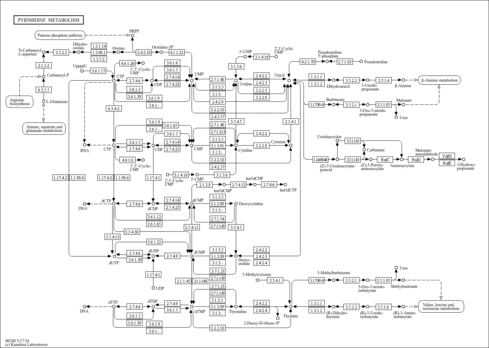
|
|---|
| Module |
| azm_M00046 | Pyrimidine degradation, uracil => beta-alanine, thymine => 3-aminoisobutanoate [PATH:azm00240] |
|
|---|
| Other DBs |
|
|---|
| Organism |
Azospirillum ramasamyi [GN: azm]
|
|---|
| Gene |
|
|---|
| Compound |
| C00119 | 5-Phospho-alpha-D-ribose 1-diphosphate |
| C00438 | N-Carbamoyl-L-aspartate |
| C00672 | 2-Deoxy-D-ribose 1-phosphate |
| C01168 | Pseudouridine 5'-phosphate |
| C01205 | (R)-3-Amino-2-methylpropanoate |
| C03997 | 5-Hydroxymethyldeoxycytidylate |
| C06198 | P1,P4-Bis(5'-uridyl) tetraphosphate |
| C11038 | 2'-Deoxy-5-hydroxymethylcytidine-5'-diphosphate |
| C11039 | 2'-Deoxy-5-hydroxymethylcytidine-5'-triphosphate |
| C15607 | 3-Oxo-3-ureidopropanoate |
| C20231 | (Z)-3-Ureidoacrylate peracid |
| C20249 | (Z)-3-Peroxyaminoacrylate |
| C20846 | 3-Oxo-3-ureidoisobutyrate |
| C21029 | (R)-3-Ureidoisobutyrate |
|
|---|
Related
pathway |
| azm00250 | Alanine, aspartate and glutamate metabolism |
| azm00280 | Valine, leucine and isoleucine degradation |
|
|---|
| KO pathway |
|
|---|
| LinkDB |
|
|---|