| Entry |
|
|---|
| Name |
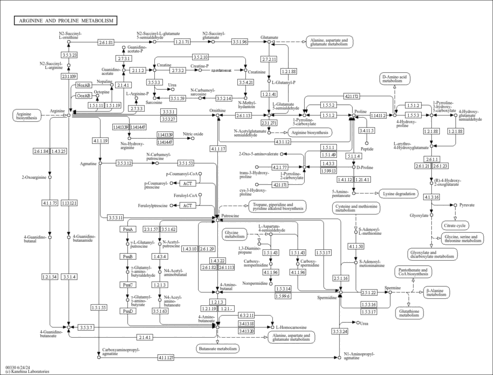
Arginine and proline metabolism - Leclercia sp. LSNIH3 |
|---|
| Class |
Metabolism; Amino acid metabolism
|
|---|
| Pathway map |
| leh00330 | Arginine and proline metabolism |
|
|---|
| Module |
| leh_M00133 | Polyamine biosynthesis, arginine => agmatine => putrescine => spermidine [PATH:leh00330] |
|
|---|
| Other DBs |
|
|---|
| Organism |
Leclercia sp. LSNIH3 [GN: leh]
|
|---|
| Gene |
|
|---|
| Compound |
| C00019 | S-Adenosyl-L-methionine |
| C00441 | L-Aspartate 4-semialdehyde |
| C01110 | 5-Amino-2-oxopentanoic acid |
| C01137 | S-Adenosylmethioninamine |
| C01165 | L-Glutamate 5-semialdehyde |
| C01250 | N-Acetyl-L-glutamate 5-semialdehyde |
| C03166 | Phosphoguanidinoacetate |
| C03415 | N2-Succinyl-L-ornithine |
| C03564 | 1-Pyrroline-2-carboxylate |
| C03771 | 5-Guanidino-2-oxopentanoate |
| C03912 | (S)-1-Pyrroline-5-carboxylate |
| C04281 | L-1-Pyrroline-3-hydroxy-5-carboxylate |
| C05147 | trans-3-Hydroxy-L-proline |
| C05932 | N-Succinyl-L-glutamate 5-semialdehyde |
| C05933 | N(omega)-Hydroxyarginine |
| C05938 | L-4-Hydroxyglutamate semialdehyde |
| C05946 | (4R)-4-Hydroxy-2-oxoglutarate |
| C05947 | L-erythro-4-Hydroxyglutamate |
| C15699 | gamma-L-Glutamylputrescine |
| C15700 | gamma-Glutamyl-gamma-aminobutyraldehyde |
| C15767 | 4-(L-gamma-Glutamylamino)butanoate |
| C19706 | cis-3-Hydroxy-L-proline |
| C20560 | N1-(3-Aminopropyl)agmatine |
| C22886 | N1-[(S)-3-amino-3-carboxypropyl]agmatine |
|
|---|
| Reference |
|
|---|
| Authors |
Haake P, Allen GW. |
|---|
| Title |
Studies on phosphorylation by phosphoroguanidinates. The mechanism of action of creatine: ATP transphosphorylase (creatine kinase). |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Kurihara S, Oda S, Kumagai H, Suzuki H. |
|---|
| Title |
Gamma-glutamyl-gamma-aminobutyrate hydrolase in the putrescine utilization pathway of Escherichia coli K-12. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Kurihara S, Oda S, Kato K, Kim HG, Koyanagi T, Kumagai H, Suzuki H. |
|---|
| Title |
A novel putrescine utilization pathway involves gamma-glutamylated intermediates of Escherichia coli K-12. |
|---|
| Journal |
|
|---|
Related
pathway |
| leh00250 | Alanine, aspartate and glutamate metabolism |
| leh00260 | Glycine, serine and threonine metabolism |
| leh00270 | Cysteine and methionine metabolism |
| leh00630 | Glyoxylate and dicarboxylate metabolism |
| leh00770 | Pantothenate and CoA biosynthesis |
|
|---|
| KO pathway |
|
|---|
| LinkDB |
|
|---|