| Entry |
|
|---|
| Name |
Glyoxylate and dicarboxylate metabolism - Sulfurifustis variabilis |
|---|
| Class |
Metabolism; Carbohydrate metabolism
|
|---|
| Pathway map |
| sva00630 | Glyoxylate and dicarboxylate metabolism |
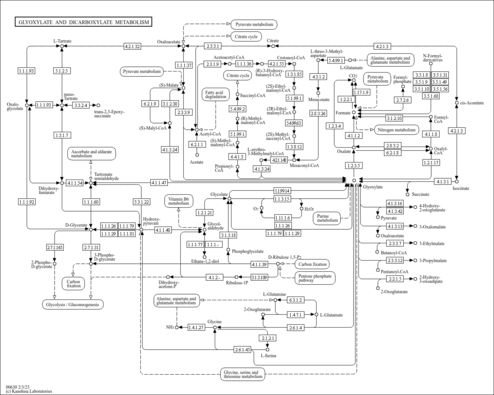
|
|---|
| Module |
|
|---|
| Other DBs |
|
|---|
| Organism |
Sulfurifustis variabilis [GN: sva]
|
|---|
| Gene |
|
|---|
| Compound |
| C01127 | 4-Hydroxy-2-oxoglutarate |
| C01146 | 2-Hydroxy-3-oxopropanoate |
| C01182 | D-Ribulose 1,5-bisphosphate |
| C03459 | 2-Hydroxy-3-oxosuccinate |
| C03548 | trans-2,3-Epoxysuccinate |
| C03561 | (R)-3-Hydroxybutanoyl-CoA |
| C03618 | L-threo-3-Methylaspartate |
| C06027 | L-erythro-3-Methylmalyl-CoA |
| C18324 | (2S)-Methylsuccinyl-CoA |
|
|---|
| Reference |
|
|---|
| Authors |
Njau RK, Herndon CA, Hawes JW. |
|---|
| Title |
Novel beta-hydroxyacid dehydrogenases in Escherichia coli and Haemophilus influenzae. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Zarzycki J, Schlichting A, Strychalsky N, Muller M, Alber BE, Fuchs G |
|---|
| Title |
Mesaconyl-coenzyme A hydratase, a new enzyme of two central carbon metabolic pathways in bacteria. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Erb TJ, Retey J, Fuchs G, Alber BE |
|---|
| Title |
Ethylmalonyl-CoA mutase from Rhodobacter sphaeroides defines a new subclade of coenzyme B12-dependent acyl-CoA mutases. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Coschigano KT, Melo-Oliveira R, Lim J, Coruzzi GM |
|---|
| Title |
Arabidopsis gls mutants and distinct Fd-GOGAT genes. Implications for photorespiration and primary nitrogen assimilation. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Masclaux-Daubresse C, Reisdorf-Cren M, Pageau K, Lelandais M, Grandjean O, Kronenberger J, Valadier MH, Feraud M, Jouglet T, Suzuki A |
|---|
| Title |
Glutamine synthetase-glutamate synthase pathway and glutamate dehydrogenase play distinct roles in the sink-source nitrogen cycle in tobacco. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Khomyakova M, Bukmez O, Thomas LK, Erb TJ, Berg IA |
|---|
| Title |
A methylaspartate cycle in haloarchaea. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Serrano JA, Bonete MJ |
|---|
| Title |
Sequencing, phylogenetic and transcriptional analysis of the glyoxylate bypass operon (ace) in the halophilic archaeon Haloferax volcanii. |
|---|
| Journal |
|
|---|
Related
pathway |
| sva00053 | Ascorbate and aldarate metabolism |
| sva00250 | Alanine, aspartate and glutamate metabolism |
| sva00260 | Glycine, serine and threonine metabolism |
| sva00710 | Carbon fixation by Calvin cycle |
|
|---|
| KO pathway |
|
|---|
| LinkDB |
|
|---|