| Entry |
|
|---|
| Name |
C5-Branched dibasic acid metabolism |
|---|
| Class |
Metabolism; Carbohydrate metabolism
|
|---|
| Pathway map |
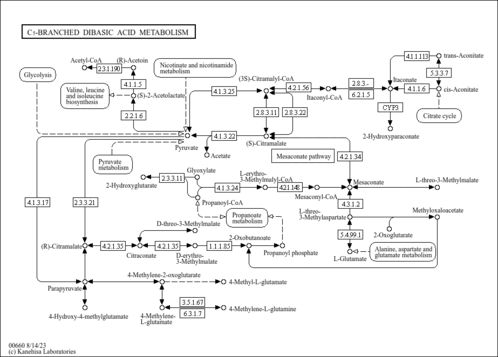
| rn00660 | C5-Branched dibasic acid metabolism |
|
|---|
| Module |
| M00535 | Isoleucine biosynthesis, pyruvate => 2-oxobutanoate [PATH:rn00660] |
|
|---|
| Reaction |
| R00008 | 4-hydroxy-4-methyl-2-oxoglutarate pyruvate-lyase (pyruvate-forming) |
| R00226 | pyruvate:pyruvate acetaldehydetransferase (decarboxylating); (S)-2-acetolactate pyruvate-lyase (carboxylating) |
| R00237 | (3S)-citramalyl-CoA pyruvate-lyase (acetyl-CoA-forming) |
| R00262 | L-threo-3-methylaspartate carboxy-aminomethylmutase |
| R00325 | (2S)-2-hydroxy-2-methylbutanedioate pyruvate-lyase (acetate-forming); (3S)-citramalate pyruvate-lyase |
| R00932 | propanoyl-CoA:glyoxylate C-propanoyltransferase (thioester-hydrolysing, 2-carboxyethyl-forming); 2-hydroxyglutarate glyoxylate-lyase (CoA-propanoylating) |
| R00934 | L-erythro-3-methylmalyl-CoA glyoxylate-lyase (propanoyl-CoA-forming) |
| R00993 | 2-oxobutanoate:oxygen 2-oxidoreductase (phosphorylating) |
| R00994 | (2R,3S)-3-methylmalate:NAD+ oxidoreductase |
| R00995 | methyloxaloacetate carboxy-lyase (2-oxobutanoate-forming) |
| R02243 | cis-aconitate carboxy-lyase |
| R02244 | trans-aconitate delta2-delta3-isomerase |
| R02404 | itaconate:CoA ligase (ADP-forming) |
| R02407 | succinyl-CoA:itaconate CoA-transferase |
| R02491 | (3S)-citramalyl-CoA hydro-lyase (itaconyl-CoA-forming) |
| R02711 | 4-methylene-L-glutamate:ammonia ligase (AMP-forming) |
| R02712 | 4-methylene-L-glutamine amidohydrolase |
| R02948 | (S)-2-hydroxy-2-methyl-3-oxobutanoate carboxy-lyase |
| R03153 | acetyl-CoA:citramalate CoA-transferase |
| R03154 | succinyl-CoA:(S)-citramalate CoA-transferase |
| R03693 | (S)-2-methylmalate hydro-lyase |
| R03696 | L-threo-3-methylaspartate ammonia-lyase |
| R03896 | (R)-2-methylmalate hydro-lyase (2-methylmaleate-forming) |
| R03898 | 2-methylmaleate hydratase; D-erythro-3-methylmalate hydro-lyase (2-methylmaleate-forming) |
| R05076 | (2R,3S)-2-methylmalyl-CoA hydro-lyase (2-methylfumaryl-CoA-forming) |
| R11749 | trans-aconitate decarboxylase |
| R13177 | acetyl-CoA:acetaldehyde acetyltransferase |
|
|---|
| Compound |
| C00651 | 4-Methylene-L-glutamate |
| C01109 | 4-Methylene-L-glutamine |
| C03618 | L-threo-3-Methylaspartate |
| C06027 | L-erythro-3-Methylmalyl-CoA |
| C06032 | D-erythro-3-Methylmalate |
| C06034 | 4-Hydroxy-4-methylglutamate |
| C06035 | 4-Methylene-2-oxoglutarate |
|
|---|
Related
pathway |
| rn00010 | Glycolysis / Gluconeogenesis |
| rn00250 | Alanine, aspartate and glutamate metabolism |
| rn00290 | Valine, leucine and isoleucine biosynthesis |
| rn00760 | Nicotinate and nicotinamide metabolism |
|
|---|
| KO pathway |
|
|---|