| Entry |
|
|---|
| Name |
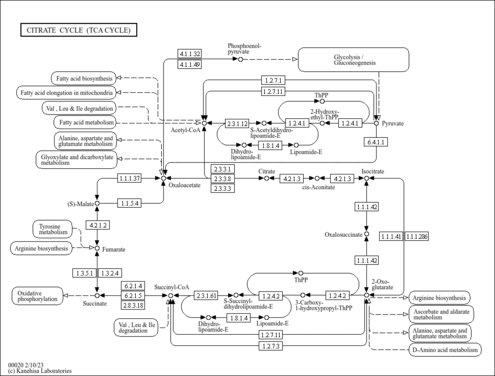
Citrate cycle (TCA cycle) |
|---|
| Description |
The citrate cycle (TCA cycle, Krebs cycle) is an important aerobic pathway for the final steps of the oxidation of carbohydrates and fatty acids. The cycle starts with acetyl-CoA, the activated form of acetate, derived from glycolysis and pyruvate oxidation for carbohydrates and from beta oxidation of fatty acids. The two-carbon acetyl group in acetyl-CoA is transferred to the four-carbon compound of oxaloacetate to form the six-carbon compound of citrate. In a series of reactions two carbons in citrate are oxidized to CO2 and the reaction pathway supplies NADH for use in the oxidative phosphorylation and other metabolic processes. The pathway also supplies important precursor metabolites including 2-oxoglutarate. At the end of the cycle the remaining four-carbon part is transformed back to oxaloacetate. According to the genome sequence data, many organisms seem to lack genes for the full cycle [MD: M00009], but contain genes for specific segments [MD: M00010 M00011].
|
|---|
| Class |
Metabolism; Carbohydrate metabolism
|
|---|
| Pathway map |
|
|---|
| Module |
| M00003 | Gluconeogenesis, oxaloacetate => fructose-6P [PATH:rn00020] |
| M00010 | Citrate cycle, first carbon oxidation, oxaloacetate => 2-oxoglutarate [PATH:rn00020] |
| M00011 | Citrate cycle, second carbon oxidation, 2-oxoglutarate => oxaloacetate [PATH:rn00020] |
|
|---|
| Other DBs |
|
|---|
| Reaction |
| R00014 | pyruvate:thiamin diphosphate acetaldehydetransferase (decarboxylating) |
| R00268 | oxalosuccinate carboxy-lyase (2-oxoglutarate-forming) |
| R00341 | ATP:oxaloacetate carboxy-lyase (transphosphorylating;phosphoenolpyruvate-forming) |
| R00342 | (S)-malate:NAD+ oxidoreductase |
| R00344 | pyruvate:carbon-dioxide ligase (ADP-forming) |
| R00351 | acetyl-CoA:oxaloacetate C-acetyltransferase (thioester-hydrolysing) |
| R00352 | acetyl-CoA:oxaloacetate C-acetyltransferase [(pro-S)-carboxymethyl-forming, ADP-phosphorylating] |
| R00361 | (S)-malate:quinone oxidoreductase |
| R00405 | succinate:CoA ligase (ADP-forming) |
| R00431 | GTP:oxaloacetate carboxy-lyase (adding GTP;phosphoenolpyruvate-forming); GTP:oxaloacetate carboxy-lyase (transphosphorylating) |
| R00432 | succinate:CoA ligase (GDP-forming) |
| R00709 | isocitrate:NAD+ oxidoreductase (decarboxylating) |
| R00726 | ITP:oxaloacetate carboxy-lyase (adding ITP; phosphoenolpyruvate-forming); ITP:oxaloacetate carboxy-lyase (transphosphorylating) |
| R00727 | succinate:CoA ligase (IDP-forming) |
| R01082 | (S)-malate hydro-lyase (fumarate-forming) |
| R01196 | pyruvate:ferredoxin 2-oxidoreductase (CoA-acetylating) |
| R01197 | 2-oxoglutarate:ferredoxin oxidoreductase (decarboxylating) |
| R01325 | citrate hydro-lyase (cis-aconitate-forming) |
| R01899 | isocitrate:NADP+ oxidoreductase |
| R01900 | isocitrate hydro-lyase (cis-aconitate-forming) |
| R02164 | succinate:quinone oxidoreductase |
| R02569 | acetyl-CoA:enzyme N6-(dihydrolipoyl)lysine S-acetyltransferase |
| R02570 | succinyl-CoA:enzyme N6-(dihydrolipoyl)lysine S-succinyltransferase |
| R07618 | enzyme N6-(dihydrolipoyl)lysine:NAD+ oxidoreductase |
| R10343 | succinyl-CoA:acetate CoA-transferase |
| R13137 | succinate:ferricytochrome-c oxidoreductase |
|
|---|
| Compound |
| C05125 | 2-(alpha-Hydroxyethyl)thiamine diphosphate |
| C05381 | 3-Carboxy-1-hydroxypropyl-ThPP |
| C15972 | Enzyme N6-(lipoyl)lysine |
| C15973 | Enzyme N6-(dihydrolipoyl)lysine |
| C16254 | [Dihydrolipoyllysine-residue succinyltransferase] S-succinyldihydrolipoyllysine |
| C16255 | [Dihydrolipoyllysine-residue acetyltransferase] S-acetyldihydrolipoyllysine |
|
|---|
| Reference |
|
|---|
| Authors |
Nishizuka Y (ed). |
|---|
| Title |
[Metabolic Maps] (In Japanese) |
|---|
| Journal |
Tokyo Kagaku Dojin (1980) |
|---|
| Reference |
|
|---|
| Authors |
Nishizuka Y, Seyama Y, Ikai A, Ishimura Y, Kawaguchi A (eds). |
|---|
| Title |
[Cellular Functions and Metabolic Maps] (In Japanese) |
|---|
| Journal |
Tokyo Kagaku Dojin (1997) |
|---|
| Reference |
|
|---|
| Authors |
Michal G. |
|---|
| Title |
Biochemical Pathways |
|---|
| Journal |
Wiley (1999) |
|---|
Related
pathway |
| rn00010 | Glycolysis / Gluconeogenesis |
| rn00053 | Ascorbate and aldarate metabolism |
| rn00250 | Alanine, aspartate and glutamate metabolism |
| rn00280 | Valine, leucine and isoleucine degradation |
| rn00630 | Glyoxylate and dicarboxylate metabolism |
|
|---|
| KO pathway |
|
|---|