 | | PATHWAY: cdet00562 | |
| Entry |
|
|---|
| Name |
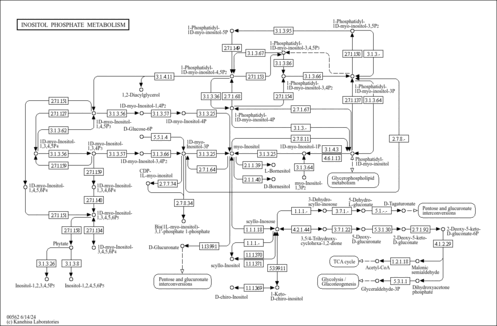
Inositol phosphate metabolism - Colletotrichum destructivum |
|---|
| Class |
Metabolism; Carbohydrate metabolism
|
|---|
| Pathway map |
|
|---|
| Other DBs |
|
|---|
| Organism |
Colletotrichum destructivum [GN: cdet]
|
|---|
| Gene |
| 87936994 | CDEST_00491; Putative phosphatidylinositol 3-/4-kinase, catalytic domain, protein kinase-like domain superfamily [KO:K19801] [EC:2.7.1.67] |
| 87937135 | CDEST_00632; Putative triosephosphate isomerase, aldolase-type TIM barrel, triosephosphate isomerase, active [KO:K01803] [EC:5.3.1.1] |
| 87939707 | CDEST_03204; Putative SAC domain, endonuclease/exonuclease/phosphatase, inositol 5-phosphatase [KO:K20279] [EC:3.1.3.36] |
| 87941087 | CDEST_04584; Putative C2 domain, phosphatidylinositol-specific phospholipase C, X domain, pleckstrin [KO:K05857] [EC:3.1.4.11] |
| 87941663 | CDEST_05160; Putative phosphatidylinositol 3-/4-kinase, catalytic domain, phosphatidylinositol 3/4-kinase [KO:K13711] [EC:2.7.1.67] |
| 87944764 | CDEST_08261; Putative C2 domain, phosphatidylinositol-specific phospholipase C, X domain, C2 domain superfamily [KO:K05857] [EC:3.1.4.11] |
| 87944801 | CDEST_08298; Putative phosphatidylinositol 3-/4-kinase, catalytic domain, protein kinase-like domain superfamily [KO:K00914] [EC:2.7.1.137] |
| 87944896 | CDEST_08393; Putative SAC domain, polyphosphoinositide phosphatase Fig4 [KO:K22913] [EC:3.1.3.-] |
| 87945120 | CDEST_08617; Putative phosphatidylinositol-specific phospholipase C, X domain-containing protein [KO:K01771] [EC:4.6.1.13] |
| 87946057 | CDEST_09554; Putative EGF-like domain, Beta propeller phytase, six-bladed beta-propeller, TolB [KO:K01083] [EC:3.1.3.8] |
| 87946434 | CDEST_09932; Putative phosphatidylinositol-specific phospholipase C, X domain-containing protein [KO:K01771] [EC:4.6.1.13] |
| 87946569 | CDEST_10067; Putative phosphatidylinositol 3-/4-kinase, catalytic domain, protein kinase-like domain superfamily [KO:K00888] [EC:2.7.1.67] |
| 87946929 | CDEST_10427; Putative triosephosphate isomerase, aldolase-type TIM barrel, triosephosphate isomerase superfamily [KO:K01803] [EC:5.3.1.1] |
| 87948611 | CDEST_12111; Putative SAC domain-containing protein [KO:K21797] [EC:3.1.3.-] |
| 87948835 | CDEST_12335; Putative FYVE zinc finger, chaperonin Cpn60/GroEL/TCP-1 family, Zinc finger, FYVE/PHD-type [KO:K00921] [EC:2.7.1.150] |
| 87950361 | CDEST_13861; Putative myo-inositol-1-phosphate synthase, NAD(P)-binding domain superfamily [KO:K01858] [EC:5.5.1.4] |
| 87951529 | CDEST_15029; Putative endonuclease/exonuclease/phosphatase, immunoglobulin-like, inositol 5-phosphatase [KO:K01099] [EC:3.1.3.36] |
|
|---|
| Compound |
| C00118 | D-Glyceraldehyde 3-phosphate |
| C00691 | 2,4,6/3,5-Pentahydroxycyclohexanone |
| C01194 | 1-Phosphatidyl-D-myo-inositol |
| C01220 | 1D-myo-Inositol 1,4-bisphosphate |
| C01243 | 1D-myo-Inositol 1,3,4-trisphosphate |
| C01245 | D-myo-Inositol 1,4,5-trisphosphate |
| C01272 | 1D-myo-Inositol 1,3,4,5-tetrakisphosphate |
| C01277 | 1-Phosphatidyl-1D-myo-inositol 4-phosphate |
| C01284 | 1D-myo-Inositol 1,3,4,5,6-pentakisphosphate |
| C03546 | myo-Inositol 4-phosphate |
| C04006 | 1D-myo-Inositol 3-phosphate |
| C04062 | D-myo-Inositol 1,3-bisphosphate |
| C04063 | D-myo-Inositol 3,4-bisphosphate |
| C04287 | 3D-3,5/4-Trihydroxycyclohexane-1,2-dione |
| C04477 | 1D-myo-Inositol 1,3,4,6-tetrakisphosphate |
| C04520 | 1D-myo-Inositol 3,4,5,6-tetrakisphosphate |
| C04549 | 1-Phosphatidyl-1D-myo-inositol 3-phosphate |
| C04563 | D-myo-Inositol 1,2,4,5,6-pentakisphosphate |
| C04579 | Inositol 1,2,3,5,6-pentakisphosphate |
| C04637 | 1-Phosphatidyl-D-myo-inositol 4,5-bisphosphate |
| C05981 | Phosphatidylinositol-3,4,5-trisphosphate |
| C06892 | 2-Deoxy-5-keto-D-gluconic acid |
| C06893 | 2-Deoxy-5-keto-D-gluconic acid 6-phosphate |
| C11174 | 1-Diphosinositol pentakisphosphate |
| C11554 | 1-Phosphatidyl-1D-myo-inositol 3,4-bisphosphate |
| C11555 | 1D-myo-Inositol 1,4,5,6-tetrakisphosphate |
| C11556 | 1-Phosphatidyl-1D-myo-inositol 3,5-bisphosphate |
| C11557 | 1-Phosphatidyl-1D-myo-inositol 5-phosphate |
| C19799 | Bis(1L-myo-inositol)-3,1'-phosphate 1-phosphate |
| C20251 | 1-Keto-D-chiro-inositol |
| C20690 | 1D-myo-Inositol 1,5-bis(diphosphate) 2,3,4,6-tetrakisphosphate |
| C21643 | 3-Dehydro-scyllo-inosose |
|
|---|
| Reference |
|
|---|
| Authors |
Yoshida K, Yamaguchi M, Ikeda H, Omae K, Tsurusaki K, Fujita Y |
|---|
| Title |
The fifth gene of the iol operon of Bacillus subtilis, iolE, encodes 2-keto-myo-inositol dehydratase. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Yoshida K, Yamaguchi M, Morinaga T, Kinehara M, Ikeuchi M, Ashida H, Fujita Y. |
|---|
| Title |
myo-Inositol catabolism in Bacillus subtilis. |
|---|
| Journal |
|
|---|
Related
pathway |
| cdet00040 | Pentose and glucuronate interconversions |
|
|---|
| KO pathway |
|
|---|
|
|