 | | PATHWAY: cdet00310 | |
| Entry |
|
|---|
| Name |
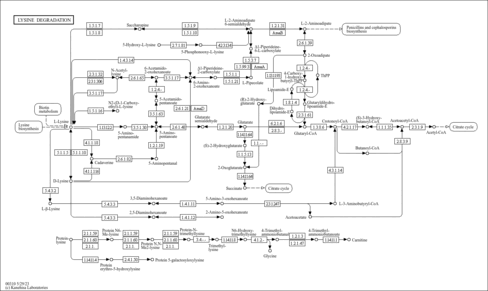
Lysine degradation - Colletotrichum destructivum |
|---|
| Class |
Metabolism; Amino acid metabolism
|
|---|
| Pathway map |
|
|---|
| Other DBs |
|
|---|
| Organism |
Colletotrichum destructivum [GN: cdet]
|
|---|
| Gene |
| 87936837 | CDEST_00334; Putative histone-lysine N-methyltransferase DOT1 domain, histone H3-K79 methyltransferase [KO:K11427] [EC:2.1.1.360] |
| 87936857 | CDEST_00354; Putative FAD dependent oxidoreductase, FAD/NAD(P)-binding domain superfamily, MTOX family [KO:K00306] [EC:1.5.3.1 1.5.3.7] |
| 87936933 | CDEST_00430; Putative enoyl-CoA hydratase/isomerase, ClpP/crotonase-like domain superfamily [KO:K07511] [EC:4.2.1.17] |
| 87937446 | CDEST_00943; Putative WW domain, SET domain, Post-SET domain, AWS domain, Set2 Rpb1 interacting [KO:K23700] |
| 87937711 | CDEST_01208; Putative enoyl-CoA hydratase/isomerase, ClpP/crotonase-like domain superfamily [KO:K07511] [EC:4.2.1.17] |
| 87938109 | CDEST_01606; Putative saccharopine dehydrogenase, NADP binding domain, NAD(P)-binding domain superfamily [KO:K00293] [EC:1.5.1.10] |
| 87938336 | CDEST_01833; Putative SET domain, histone-lysine N-methyltransferase domain, SET domain superfamily [KO:K11429] [EC:2.1.1.362] |
| 87938927 | CDEST_02424; Putative FAD dependent oxidoreductase, FAD/NAD(P)-binding domain superfamily, MTOX family [KO:K00306] [EC:1.5.3.1 1.5.3.7] |
| 87938970 | CDEST_02467; Putative biotin/lipoyl attachment, 2-oxoacid dehydrogenase acyltransferase, catalytic [KO:K00658] [EC:2.3.1.61] |
| 87939850 | CDEST_03347; Putative biotin/lipoyl attachment, 2-oxoacid dehydrogenase acyltransferase, catalytic [KO:K00658] [EC:2.3.1.61] |
| 87939878 | CDEST_03375; Putative pyridine nucleotide-disulfide oxidoreductase, class I, FAD/NAD(P)-binding protein [KO:K00382] [EC:1.8.1.4] |
| 87941158 | CDEST_04655; Putative TauD/TfdA-like domain, gamma-butyrobetaine hydroxylase-like, trimethyllysine dioxygenase [KO:K00474] [EC:1.14.11.8] |
| 87941438 | CDEST_04935; Putative SET domain, nucleotide-binding alpha-beta plait domain superfamily, SET domain superfamily [KO:K11422] [EC:2.1.1.354] |
| 87943428 | CDEST_06925; Putative aldehyde dehydrogenase domain, aldehyde/histidinol dehydrogenase [KO:K00128] [EC:1.2.1.3] |
| 87943635 | CDEST_07132; Putative FAD dependent oxidoreductase, FAD/NAD(P)-binding domain superfamily, MTOX family [KO:K00306] [EC:1.5.3.1 1.5.3.7] |
| 87943785 | CDEST_07282; Putative FAD dependent oxidoreductase, FAD/NAD(P)-binding domain superfamily, MTOX family [KO:K00306] [EC:1.5.3.1 1.5.3.7] |
| 87943860 | CDEST_07357; Putative aldehyde dehydrogenase domain, aldehyde/histidinol dehydrogenase [KO:K00128] [EC:1.2.1.3] |
| 87945918 | CDEST_09415; Putative aldehyde dehydrogenase domain, aldehyde/histidinol dehydrogenase [KO:K00128] [EC:1.2.1.3] |
| 87946946 | CDEST_10444; Putative acyl-CoA oxidase/dehydrogenase, middle domain, acyl-CoA dehydrogenase/oxidase [KO:K00252] [EC:1.3.8.6] |
| 87947628 | CDEST_11128; Putative FAD dependent oxidoreductase, FAD/NAD(P)-binding domain superfamily, MTOX family [KO:K00306] [EC:1.5.3.1 1.5.3.7] |
| 87948791 | CDEST_12291; Putative alanine dehydrogenase/pyridine nucleotide transhydrogenase, NAD(H)-binding protein [KO:K00290] [EC:1.5.1.7] |
| 87949587 | CDEST_13087; Putative FAD dependent oxidoreductase, FAD/NAD(P)-binding domain superfamily, MTOX family [KO:K00306] [EC:1.5.3.1 1.5.3.7] |
| 87949984 | CDEST_13484; Putative aldehyde dehydrogenase domain, aldehyde/histidinol dehydrogenase [KO:K00128] [EC:1.2.1.3] |
| 87950038 | CDEST_13538; Putative saccharopine dehydrogenase, NADP binding domain, NAD(P)-binding domain superfamily [KO:K00293] [EC:1.5.1.10] |
| 87950124 | CDEST_13624; Putative FAD dependent oxidoreductase, FAD/NAD(P)-binding domain superfamily, MTOX family [KO:K00306] [EC:1.5.3.1 1.5.3.7] |
|
|---|
| Compound |
| C00449 | N6-(L-1,3-Dicarboxypropyl)-L-lysine |
| C00450 | (S)-2,3,4,5-Tetrahydropyridine-2-carboxylate |
| C01142 | (3S)-3,6-Diaminohexanoate |
| C01144 | (S)-3-Hydroxybutanoyl-CoA |
| C01149 | 4-Trimethylammoniobutanal |
| C01181 | 4-Trimethylammoniobutanoate |
| C01186 | (3S,5S)-3,5-Diaminohexanoate |
| C01211 | Procollagen 5-hydroxy-L-lysine |
| C01259 | (3S)-3-Hydroxy-N6,N6,N6-trimethyl-L-lysine |
| C03656 | (S)-5-Amino-3-oxohexanoic acid |
| C03793 | N6,N6,N6-Trimethyl-L-lysine |
| C04076 | L-2-Aminoadipate 6-semialdehyde |
| C04092 | Delta1-Piperideine-2-carboxylate |
| C04487 | 5-(D-Galactosyloxy)-L-lysine-procollagen |
| C05161 | (2R,5S)-2,5-Diaminohexanoate |
| C05544 | Protein N6-methyl-L-lysine |
| C05545 | Protein N6,N6-dimethyl-L-lysine |
| C05546 | Protein N6,N6,N6-trimethyl-L-lysine |
| C05548 | 6-Acetamido-2-oxohexanoate |
| C06157 | [Dihydrolipoyllysine-residue succinyltransferase] S-glutaryldihydrolipoyllysine |
| C15972 | Enzyme N6-(lipoyl)lysine |
| C15973 | Enzyme N6-(dihydrolipoyl)lysine |
| C22667 | 4-Carboxy-1-hydroxybutyryl-ThPP |
|
|---|
| Reference |
|
|---|
| Authors |
Vaz FM, Wanders RJ. |
|---|
| Title |
Carnitine biosynthesis in mammals. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Large PJ, Robertson A. |
|---|
| Title |
The route of lysine breakdown in Candida tropicalis. |
|---|
| Journal |
|
|---|
Related
pathway |
| cdet00311 | Penicillin and cephalosporin biosynthesis |
|
|---|
| KO pathway |
|
|---|
|
|