| Entry |
|
|---|
| Name |
N6-(L-1,3-dicarboxypropyl)-L-lysine:NAD+ oxidoreductase (L-lysine-forming)
|
|---|
| Definition |
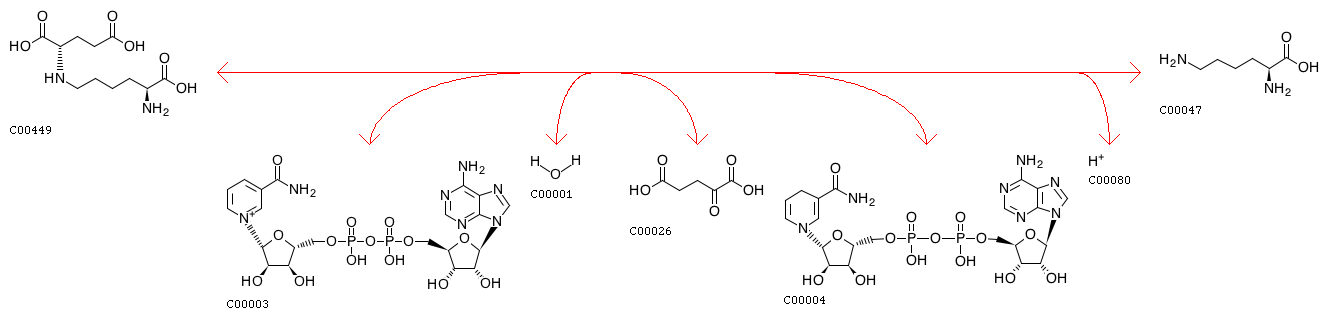
N6-(L-1,3-Dicarboxypropyl)-L-lysine + NAD+ + H2O <=> L-Lysine + 2-Oxoglutarate + NADH + H+
|
|---|
| Equation |
|
|---|
|
 |
|---|
| Reaction class |
|
|---|
| Enzyme |
|
|---|
| Pathway |
| rn01110 | Biosynthesis of secondary metabolites |
| rn01230 | Biosynthesis of amino acids |
|
|---|
| Module |
| M00030 | Lysine biosynthesis, AAA pathway, 2-oxoglutarate => 2-aminoadipate => lysine |
|
|---|
| Brite |
Enzymatic reactions [BR:br08201]
1. Oxidoreductase reactions
1.5 Acting on the CH-NH group of donors
1.5.1 With NAD+ or NADP+ as acceptor
1.5.1.7
R00715 N6-(L-1,3-Dicarboxypropyl)-L-lysine + NAD+ + H2O <=> L-Lysine + 2-Oxoglutarate + NADH + H+
|
|---|
| Orthology |
| K00290 | saccharopine dehydrogenase (NAD+, L-lysine forming) [EC:1.5.1.7] |
|
|---|
| Other DBs |
|
|---|
| LinkDB |
|
|---|