| Entry |
|
|---|
| Name |
Antigen processing and presentation - Herpailurus yagouaroundi (jaguarundi) |
|---|
| Class |
Organismal Systems; Immune system
|
|---|
| Pathway map |
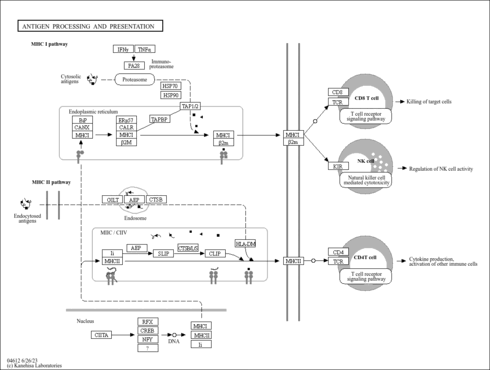
| pyu04612 | Antigen processing and presentation |
|
|---|
| Other DBs |
|
|---|
| Organism |
Herpailurus yagouaroundi (jaguarundi) [GN: pyu]
|
|---|
| Gene |
| 121011003 | IFI30; gamma-interferon-inducible lysosomal thiol reductase [KO:K08059] |
| 121026934 | NFYC; nuclear transcription factor Y subunit gamma isoform X1 [KO:K08066] |
| 121029419 | NFYA; nuclear transcription factor Y subunit alpha isoform X1 [KO:K08064] |
| 121030974 | CREB1; cyclic AMP-responsive element-binding protein 1 isoform X1 [KO:K05870] |
| 121040510 | NFYB; nuclear transcription factor Y subunit beta isoform X1 [KO:K08065] |
| 121044489 | CD8A; T-cell surface glycoprotein CD8 alpha chain isoform X1 [KO:K06458] |
| 121045050 | PSME3; proteasome activator complex subunit 3 isoform X1 [KO:K06698] |
| 121045086 | CD74; HLA class II histocompatibility antigen gamma chain isoform X1 [KO:K06505] |
|
|---|
| Reference |
|
|---|
| Authors |
Trombetta ES, Mellman I. |
|---|
| Title |
Cell biology of antigen processing in vitro and in vivo. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Inaba K, Inaba M. |
|---|
| Title |
Antigen recognition and presentation by dendritic cells. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Heath WR, Carbone FR. |
|---|
| Title |
Cross-presentation in viral immunity and self-tolerance. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Abele R, Tampe R. |
|---|
| Title |
The ABCs of immunology: structure and function of TAP, the transporter associated with antigen processing. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Reith W, LeibundGut-Landmann S, Waldburger JM. |
|---|
| Title |
Regulation of MHC class II gene expression by the class II transactivator. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Honey K, Rudensky AY. |
|---|
| Title |
Lysosomal cysteine proteases regulate antigen presentation. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Sercarz EE, Maverakis E. |
|---|
| Title |
Mhc-guided processing: binding of large antigen fragments. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Ackerman AL, Cresswell P. |
|---|
| Title |
Cellular mechanisms governing cross-presentation of exogenous antigens. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Cresswell P, Ackerman AL, Giodini A, Peaper DR, Wearsch PA. |
|---|
| Title |
Mechanisms of MHC class I-restricted antigen processing and cross-presentation. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Zika E, Ting JP. |
|---|
| Title |
Epigenetic control of MHC-II: interplay between CIITA and histone-modifying enzymes. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Reith W, Mach B. |
|---|
| Title |
The bare lymphocyte syndrome and the regulation of MHC expression. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Lizee G, Basha G, Jefferies WA. |
|---|
| Title |
Tails of wonder: endocytic-sorting motifs key for exogenous antigen presentation. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Watts C. |
|---|
| Title |
The exogenous pathway for antigen presentation on major histocompatibility complex class II and CD1 molecules. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Kaufmann SH, Schaible UE. |
|---|
| Title |
Antigen presentation and recognition in bacterial infections. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Groettrup M, Kirk CJ, Basler M |
|---|
| Title |
Proteasomes in immune cells: more than peptide producers? |
|---|
| Journal |
|
|---|
Related
pathway |
| pyu04650 | Natural killer cell mediated cytotoxicity |
| pyu04660 | T cell receptor signaling pathway |
|
|---|
| KO pathway |
|
|---|