| Entry |
|
|---|
| Name |
Tyrosine metabolism - Gorilla gorilla gorilla (western lowland gorilla) |
|---|
| Class |
Metabolism; Amino acid metabolism
|
|---|
| Pathway map |
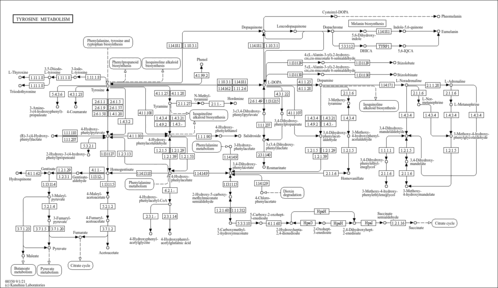
|
|---|
| Module |
| ggo_M00042 | Catecholamine biosynthesis, tyrosine => dopamine => noradrenaline => adrenaline [PATH:ggo00350] |
| ggo_M00043 | Thyroid hormone biosynthesis, tyrosine => triiodothyronine/thyroxine [PATH:ggo00350] |
|
|---|
| Other DBs |
|
|---|
| Organism |
Gorilla gorilla gorilla (western lowland gorilla) [GN: ggo]
|
|---|
| Gene |
| 101138123 | TYRP1; 5,6-dihydroxyindole-2-carboxylic acid oxidase [KO:K00506] [EC:1.14.18.-] |
|
|---|
| Compound |
| C00355 | 3,4-Dihydroxy-L-phenylalanine |
| C01161 | 3,4-Dihydroxyphenylacetate |
| C01179 | 3-(4-Hydroxyphenyl)pyruvate |
| C03765 | 4-Hydroxyphenylacetaldehyde |
| C03964 | (R)-3-(4-Hydroxyphenyl)lactate |
| C04043 | 3,4-Dihydroxyphenylacetaldehyde |
| C04045 | 3-(3,4-Dihydroxyphenyl)pyruvate |
| C04052 | 5-Carboxy-2-oxohept-3-enedioate |
| C04185 | 5,6-Dihydroxyindole-2-carboxylate |
| C04186 | 5-Carboxymethyl-2-hydroxymuconate |
| C04368 | 3-Amino-3-(4-hydroxyphenyl)propanoate |
| C04642 | 2-Hydroxy-5-carboxymethylmuconate semialdehyde |
| C04796 | 4-(L-Alanin-3-yl)-2-hydroxy-cis,cis-muconate 6-semialdehyde |
| C04797 | 5-(L-Alanin-3-yl)-2-hydroxy-cis,cis-muconate 6-semialdehyde |
| C05338 | 4-Hydroxyphenylacetyl-CoA |
| C05350 | 2-Hydroxy-3-(4-hydroxyphenyl)propenoate |
| C05576 | 3,4-Dihydroxyphenylethyleneglycol |
| C05577 | 3,4-Dihydroxymandelaldehyde |
| C05581 | 3-Methoxy-4-hydroxyphenylacetaldehyde |
| C05583 | 3-Methoxy-4-hydroxyphenylglycolaldehyde |
| C05584 | 3-Methoxy-4-hydroxymandelate |
| C05594 | 3-Methoxy-4-hydroxyphenylethyleneglycol |
| C05595 | 4-Hydroxyphenylacetylglutamic acid |
| C05596 | p-Hydroxyphenylacetylglycine |
| C05600 | 2-Hydroxyhepta-2,4-dienedioate |
| C05604 | 2-Carboxy-2,3-dihydro-5,6-dihydroxyindole |
| C06201 | 2,4-Dihydroxyhept-2-enedioate |
| C10447 | 3,4-Dihydroxyphenylpropanoate |
| C17938 | 5,6-Indolequinone-2-carboxylic acid |
| C22038 | (R)-3-(3,4-Dihydroxyphenyl)lactate |
|
|---|
| Reference |
|
|---|
| Authors |
Prieto MA, Diaz E, Garcia JL. |
|---|
| Title |
Molecular characterization of the 4-hydroxyphenylacetate catabolic pathway of Escherichia coli W: engineering a mobile aromatic degradative cluster. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Sparnins VL, Dagley S. |
|---|
| Title |
Alternative routes of aromatic catabolism in Pseudomonas acidovorans and Pseudomonas putida: gallic acid as a substrate and inhibitor of dioxygenases. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Ito S, Wakamatsu K |
|---|
| Title |
Chemistry of mixed melanogenesis--pivotal roles of dopaquinone. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Olivares C, Jimenez-Cervantes C, Lozano JA, Solano F, Garcia-Borron JC |
|---|
| Title |
The 5,6-dihydroxyindole-2-carboxylic acid (DHICA) oxidase activity of human tyrosinase. |
|---|
| Journal |
|
|---|
Related
pathway |
| ggo00400 | Phenylalanine, tyrosine and tryptophan biosynthesis |
|
|---|
| KO pathway |
|
|---|