| Entry |
|
|---|
| Name |
ATP-dependent chromatin remodeling - Ictalurus furcatus (blue catfish) |
|---|
| Class |
Genetic Information Processing; Chromosome
|
|---|
| Pathway map |
| ifu03082 | ATP-dependent chromatin remodeling |
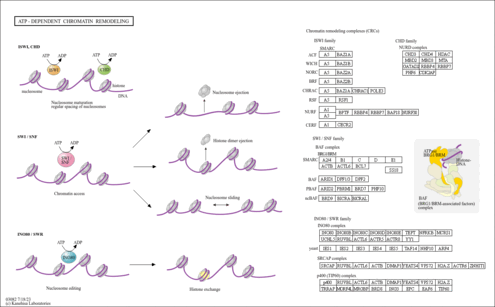
|
|---|
| Organism |
Ictalurus furcatus (blue catfish) [GN: ifu]
|
|---|
| Gene |
| 128606059 | smarca5; SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily A member 5 [KO:K11654] [EC:5.6.2.-] |
| 128612260 | baz1a; bromodomain adjacent to zinc finger domain protein 1A isoform X1 [KO:K11655] |
| 128619885 | baz2a; bromodomain adjacent to zinc finger domain protein 2A isoform X1 [KO:K15224] |
| 128608932 | bromodomain adjacent to zinc finger domain protein 2B isoform X1 [KO:K26167] |
| 128615678 | baz2ba; bromodomain adjacent to zinc finger domain protein 2B isoform X1 [KO:K26167] |
| 128603770 | smarca1; probable global transcription activator SNF2L1 isoform X1 [KO:K11727] [EC:5.6.2.-] |
| 128615388 | nucleosome-remodeling factor subunit BPTF-like isoform X1 [KO:K11728] |
| 128599112 | bptf; nucleosome-remodeling factor subunit BPTF isoform X1 [KO:K11728] |
| 128620245 | c16h17orf49; chromatin complexes subunit BAP18 isoform X1 [KO:K23406] |
| 128610050 | zgc:92664; chromatin complexes subunit BAP18 isoform X1 [KO:K23406] |
| 128610083 | chd3; chromodomain-helicase-DNA-binding protein 3 isoform X1 [KO:K11642] [EC:5.6.2.-] |
| 128617426 | chd4b; chromodomain-helicase-DNA-binding protein 4 [KO:K11643] [EC:5.6.2.-] |
| 128615707 | chd4a; chromodomain-helicase-DNA-binding protein 4a [KO:K11643] [EC:5.6.2.-] |
| 128613455 | mbd3b; methyl-CpG-binding domain protein 3b isoform X1 [KO:K11591] |
| 128624387 | mbd3a; methyl-CpG-binding domain protein 3a isoform X1 [KO:K11591] |
| 128613666 | gatad2ab; GATA zinc finger domain containing 2Ab isoform X1 [KO:K23194] |
| 128600346 | gatad2b; transcriptional repressor p66-beta isoform X1 [KO:K23194] |
| 128620163 | cyclin-dependent kinase 2-associated protein 1 isoform X1 [KO:K26454] |
| 128606138 | cdk2ap2; cyclin-dependent kinase 2-associated protein 2 [KO:K26455] |
| 128599545 | smarca4a; SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 4a isoform X1 [KO:K11647] [EC:5.6.2.-] |
| 128625589 | smarca2; probable global transcription activator SNF2L2 isoform X1 [KO:K11647] [EC:5.6.2.-] |
| 128622534 | SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily B member 1-A isoform X1 [KO:K11648] |
| 128600622 | smarcc1b; SWI/SNF complex subunit SMARCC1b isoform X1 [KO:K11649] |
| 128617515 | smarcd2; SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily D member 2 [KO:K11650] |
| 128609562 | smarcd3a; SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily D member 3 isoform X1 [KO:K11650] |
| 128609623 | smarcd3b; SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily D member 3b isoform X1 [KO:K11650] |
| 128625135 | smarcd1; SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily D member 1 [KO:K11650] |
| 128622004 | smarce1; SWI/SNF-related matrix-associated actin-dependent regulator of chromatin subfamily E member 1 isoform X1 [KO:K11651] |
| 128620044 | bcl7ba; B-cell CLL/lymphoma 7 protein family member B-A [KO:K25605] |
| 128606965 | bcl7bb; B-cell CLL/lymphoma 7 protein family member B-B [KO:K25605] |
| 128598963 | bcl7a; B-cell CLL/lymphoma 7 protein family member A isoform X1 [KO:K25605] |
| 128612572 | arid1aa; AT-rich interactive domain-containing protein 1A isoform X1 [KO:K11653] |
| 128612729 | AT-rich interactive domain-containing protein 1B-like [KO:K11653] |
| 128616176 | arid1ab; AT-rich interactive domain-containing protein 1A [KO:K11653] |
| 128601205 | arid1b; AT-rich interactive domain-containing protein 1B isoform X1 [KO:K11653] |
| 128617691 | arid2; AT-rich interactive domain-containing protein 2 isoform X1 [KO:K11765] |
| 128606807 | bicra; BRD4-interacting chromatin-remodeling complex-associated protein isoform X1 [KO:K25612] |
| 128617379 | bicral; BRD4-interacting chromatin-remodeling complex-associated protein-like [KO:K25613] |
| 128600915 | ino80; chromatin-remodeling ATPase INO80 isoform X1 [KO:K11665] [EC:5.6.2.-] |
| 128622454 | nfrkb; nuclear factor related to kappa-B-binding protein isoform X1 [KO:K11671] |
| 128601038 | yy1b; transcriptional repressor protein YY1b isoform X1 [KO:K09201] |
| 128624772 | vps72a; vacuolar protein sorting 72 homolog a isoform X1 [KO:K11664] |
| 128607391 | ep400; E1A-binding protein p400 isoform X1 [KO:K11320] [EC:5.6.2.-] |
| 128616534 | trrap; transformation/transcription domain-associated protein isoform X1 [KO:K08874] |
|
|---|
| Reference |
|
|---|
| Authors |
Hasan N, Ahuja N |
|---|
| Title |
The Emerging Roles of ATP-Dependent Chromatin Remodeling Complexes in Pancreatic Cancer. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
He S, Wu Z, Tian Y, Yu Z, Yu J, Wang X, Li J, Liu B, Xu Y |
|---|
| Title |
Structure of nucleosome-bound human BAF complex. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Wanior M, Kramer A, Knapp S, Joerger AC |
|---|
| Title |
Exploiting vulnerabilities of SWI/SNF chromatin remodelling complexes for cancer therapy. |
|---|
| Journal |
|
|---|
| Reference |
|
|---|
| Authors |
Burgio G, Onorati MC, Corona DF |
|---|
| Title |
Chromatin remodeling regulation by small molecules and metabolites. |
|---|
| Journal |
|
|---|
| KO pathway |
|
|---|
| LinkDB |
|
|---|