| Entry |
|
|---|
| Definition |
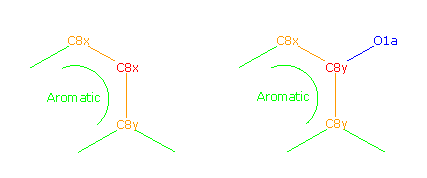
C8x-C8y:*-O1a:C8x+C8y-C8x+C8y
|
|---|
|
 |
|---|
| Reactant pair |
|
|---|
| Related class |
|
|---|
| Reaction |
|
|---|
| Enzyme |
|
|---|
| Pathway |
| rn00130 | Ubiquinone and other terpenoid-quinone biosynthesis |
| rn00140 | Steroid hormone biosynthesis |
| rn00361 | Chlorocyclohexane and chlorobenzene degradation |
| rn00624 | Polycyclic aromatic hydrocarbon degradation |
| rn00760 | Nicotinate and nicotinamide metabolism |
| rn00901 | Indole alkaloid biosynthesis |
| rn00940 | Phenylpropanoid biosynthesis |
| rn00944 | Flavone and flavonol biosynthesis |
| rn00945 | Stilbenoid, diarylheptanoid and gingerol biosynthesis |
| rn00950 | Isoquinoline alkaloid biosynthesis |
| rn00980 | Metabolism of xenobiotics by cytochrome P450 |
| rn00982 | Drug metabolism - cytochrome P450 |
| rn00996 | Biosynthesis of various alkaloids |
| rn00999 | Biosynthesis of various plant secondary metabolites |
| rn01057 | Biosynthesis of type II polyketide products |
| rn01059 | Biosynthesis of enediyne antibiotics |
| rn01110 | Biosynthesis of secondary metabolites |
| rn01120 | Microbial metabolism in diverse environments |
| rn01220 | Degradation of aromatic compounds |
|
|---|
| RModule |
| RM006 | Dihydroxylation of aromatic ring, type 2 (two monooxygenase reactions) |
| RM007 | Dihydroxylation of aromatic ring, type 3 (dealkylation and monooxygenase reactions) |
| RM012 | Dihydroxylation and meta-cleavage of aromatic ring, type 3a |
| RM026 | Hydoxylation and decarboxylation motif |
| RM027 | Hydroxylation and methylation motif |
| RM029 | Pterocarpan synthesis |
|
|---|
| Orthology |
| K09755 | ferulate-5-hydroxylase [EC:1.14.-.-] |
| K13267 | flavonoid 6-hydroxylase [EC:1.14.13.-] |
| K14630 | two-component flavin-dependent monooxygenase [EC:1.14.14.-] |
| K15930 | bifunctional oxygenase/reductase |
| K16249 | phenol/toluene 2-monooxygenase (NADH) P0/A0 |
| K21727 | 4-nitrophenol 2-monooxygenase / 4-nitrocatechol 4-monooxygenase, reductase component |
| K23470 | 4-hydroxyphenylacetate 3-hydroxylase, reductase component [EC:1.5.1.36] |
| K24844 | 2-(all-trans-polyprenyl)phenol 6-hydroxylase (prephenate) UbiU [EC:1.97.1.15] |
| K24845 | 2-(all-trans-polyprenyl)phenol 6-hydroxylase (prephenate) UbiV [EC:1.97.1.15] |
| K26037 | tyrosine hydroxylase / L-DOPA oxygenase [EC:1.14.-.-] |
|
|---|